Description
Babson Analytics and Quantitative Methods Tools.
Description
Instructor-developed tools for Analytics and Quantitative Methods (AQM) courses at Babson College. Included are compact descriptive statistics for data frames and lists, expanded reporting and graphics for linear regressions, and formatted reports for best subsets analyses.
README.md
BAQM
BAQM supplies functions developed by Babson College instructors for AQM 1000 and AQM 2000 courses using R in the curriculum. The primary functions provide:
sumry.df()- summary descriptive statistics for data frames, allowing both numeric and factor variables, in both wide and long formats.sumry.lm()- expanded statistics and enhanced formatting to summarize linear model results.lm_plot.4way()- multiple diagnostic plots for linear models using ggplot, including a 4-in-1 summary graphic.sumry.regsubsets()- compact “best subsets” linear model reports for analytics from theregsubsetsfunction of the leaps package.
Installation
You can install the development version of BAQM from GitHub with:
install.packages("pak")
pak::pak("CPA-wrk/BAQM")
Example
These examples use the built-in R data sets iris, swiss, and mtcars, and show:
- brief statistical summaries of data sets
irisandswiss, - a concise best-subsets report for regression modeling of Fertility in
swiss, - expanded statistical summaries for linear models of Sepal Length in
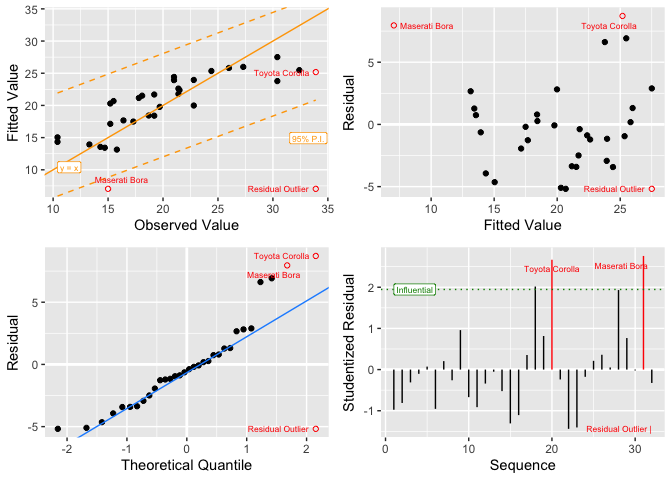
irisand miles per gallon inmtcars, and - a 4-way diagnostic plot for the
mtcarsmodel.
(Variable names are truncated in swiss to narrow the output.)
library(leaps)
library(BAQM)
#
sumry(iris) # Includes non-numeric variable
#> Sepal.Length Sepal.Width Petal.Length Petal.Width Species
#> n.val 150 150 150 150 150
#> n.na 0 0 0 0 0
#> min 4.3 2 1 0.1 n.lvl : 3
#> Q1 5.1 2.7 1.6 0.2 setosa :50
#> median 5.8 3 4.35 1.3 versclr:50
#> mean 5.843 3.057 3.758 1.199 virginc:50
#> Q3 6.45 3.4 5.1 1.8
#> max 7.9 4.4 6.9 2.5
#> std.dev 0.8281 0.4359 1.765 0.7622
#
names(swiss) # Show original variable names
#> [1] "Fertility" "Agriculture" "Examination" "Education"
#> [5] "Catholic" "Infant.Mortality"
names(swiss) <- substr(names(swiss), 1, 4) # Narrows output
sumry(swiss)
#> Fert Agri Exam Educ Cath Infa
#> n.val 47 47 47 47 47 47
#> n.na 0 0 0 0 0 0
#> min 35 1.2 3 1 2.15 10.8
#> Q1 64.4 35.3 12 6 5.16 18
#> median 70.4 54.1 16 8 15.14 20
#> mean 70.14 50.66 16.49 10.98 41.14 19.94
#> Q3 79.3 67.8 22 12.5 93.4 22.2
#> max 92.5 89.7 37 53 100 26.6
#> std.dev 12.49 22.71 7.978 9.615 41.7 2.913
regs <- regsubsets(Fert ~ ., data = swiss, nbest = 3)
sumry(regs)
#>
#> Call: (function (...)
#> rmarkdown::render(...))(input = base::quote("/Users/peter/Library/CloudStorage/OneDrive-centerpointanalytics.com/CPA_wrk/R/BAQM/README.Rmd"),
#> output_options = base::quote(list(html_preview = FALSE)),
#> quiet = base::quote(TRUE))
#> _k_i.best rsq adjr2 see cp Agri Exam Educ Cath Infa
#> 1 1 ( 1 ) 0.4406 0.4282 9.446029 35.20 *
#> 2 1 ( 2 ) 0.4172 0.4042 9.642000 38.48 *
#> 3 1 ( 3 ) 0.2150 0.1976 11.189945 66.75 *
#> 4 2 ( 1 ) 0.5745 0.5552 8.331442 18.49 * *
#> 5 2 ( 2 ) 0.5648 0.5450 8.426136 19.85 * *
#> 6 2 ( 3 ) 0.5363 0.5152 8.697447 23.83 * *
#> 7 3 ( 1 ) 0.6625 0.6390 7.505417 8.18 * * *
#> 8 3 ( 2 ) 0.6423 0.6173 7.727757 11.01 * * *
#> 9 3 ( 3 ) 0.6191 0.5925 7.973957 14.25 * * *
#> 10 4 ( 1 ) 0.6993 0.6707 7.168166 5.03 * * * *
#> 11 4 ( 2 ) 0.6639 0.6319 7.579356 9.99 * * * *
#> 12 4 ( 3 ) 0.6498 0.6164 7.736422 11.96 * * * *
#> 13 5 ( 1 ) 0.7067 0.6710 7.165369 6.00 * * * * *
#
mdl <- lm(Sepal.Length ~ ., data = iris)
sumry(mdl)
#>
#> Summary Statistics:
#> Value Performance Measure Err(Resids) Metric
#> Observations = 150 R-Squared = 0.86731 MAPE = 0.041785
#> F-Statistic = 188.25 Adj-R2 = 0.86271 MAD = 0.24286
#> Pr(b's=0) = <2e-16 *** Std.Err.Est = 0.30683 RMSE = 0.30063
#>
#> Analysis of Variance:
#> Deg.Frdm Sum.of.Sqs Mean.Sum.Sqs F.statistic p-value(F)
#> Regression 5 88.612 17.722370 188.25 <2e-16 ***
#> Error(Resids) 144 13.556 0.094142
#> Total 149 102.168
#>
#> Coefficients:
#> Coefficient Std.Error t-stat p-value VIF
#> (Intercept) 2.17127 0.279794 7.7602 1.43e-12 ***
#> Sepal.Width 0.49589 0.086070 5.7615 4.87e-08 *** 2.2275
#> Petal.Length 0.82924 0.068528 12.1009 < 2e-16 *** 23.1616
#> Petal.Width -0.31516 0.151196 -2.0844 0.03889 * 21.0214
#> Species_versicolor -0.72356 0.240169 -3.0127 0.00306 ** 20.4234
#> Species_virginica -1.02350 0.333726 -3.0669 0.00258 ** 39.4344
#>
#> Signif.Levels: 0 '***' 0.001 '** ' 0.01 ' * ' 0.05 ' . ' 0.1 ' ' 1
#>
#> Summary of Min 1Q Mean Median 3Q Max
#> Residuals: -0.7942 -0.2187 <3e-14 0.008987 0.2025 0.731
#>
#> Call: lm(formula = Sepal.Length ~ ., data = iris)
#
mdl <- lm(mpg ~ hp + qsec, data = mtcars)
sumry(mdl)
#>
#> Summary Statistics:
#> Value Performance Measure Err(Resids) Metric
#> Observations = 32 R-Squared = 0.63688 MAPE = 0.1448
#> F-Statistic = 25.431 Adj-R2 = 0.61183 MAD = 2.6984
#> Pr(b's=0) = 4.18e-07 *** Std.Err.Est = 3.755 RMSE = 3.5746
#>
#> Analysis of Variance:
#> Deg.Frdm Sum.of.Sqs Mean.Sum.Sqs F.statistic p-value(F)
#> Regression 2 717.15 358.58 25.431 4.18e-07 ***
#> Error(Resids) 29 408.89 14.10
#> Total 31 1126.05
#>
#> Coefficients:
#> Coefficient Std.Error t-stat p-value VIF
#> (Intercept) 48.323705 11.103306 4.3522 0.000153 ***
#> hp -0.084593 0.013933 -6.0715 1.31e-06 *** 2.0063
#> qsec -0.886580 0.534585 -1.6584 0.108007 2.0063
#>
#> Signif.Levels: 0 '***' 0.001 '** ' 0.01 ' * ' 0.05 ' . ' 0.1 ' ' 1
#>
#> Summary of Min 1Q Median Mean 3Q Max
#> Residuals: -5.178 -2.603 -0.5098 <6e-15 1.287 8.718
#>
#> Call: lm(formula = mpg ~ hp + qsec, data = mtcars)
lm_plot.4way(mdl)