Description
Nested Sequential Monte Carlo for the Bayesian Mallows Model.
Description
Provides nested sequential Monte Carlo algorithms for performing sequential inference in the Bayesian Mallows model, which is a widely used probability model for rank and preference data. The package implements the SMC2 (Sequential Monte Carlo Squared) algorithm for handling sequentially arriving rankings and pairwise preferences, including support for complete rankings, partial rankings, and pairwise comparisons. The methods are based on Sorensen (2025) <doi:10.1214/25-BA1564>.
README.md
BayesMallowsSMC2
BayesMallowsSMC2 provides functions for performing sequential inference in the Bayesian Mallows model using the SMC2 algorithm (Sørensen et al. 2025).
Installation
You can install the development version of BayesMallowsSMC2 from GitHub with:
# install.packages("devtools")
devtools::install_github("osorensen/BayesMallowsSMC2")
Example
Here is a basic example using the included complete_rankings dataset:
library(BayesMallowsSMC2)
# Fit the model with complete rankings
set.seed(123)
mod <- compute_sequentially(
complete_rankings,
hyperparameters = set_hyperparameters(n_items = 5),
smc_options = set_smc_options(n_particles = 100, n_particle_filters = 1)
)
# Show model info
mod
#> BayesMallowsSMC2 Model
#> ======================
#>
#> Number of particles: 100
#> Number of timepoints: 100
#> Number of items: 5
#> Number of clusters: 1
#>
#> Log marginal likelihood: -472.34
#> Final ESS: 63.55
#> Resampling events: 5/100
Posterior Summaries
We can visualize the posterior distributions of the parameters:
# Posterior distribution of alpha (dispersion parameter)
plot(mod, parameter = "alpha")
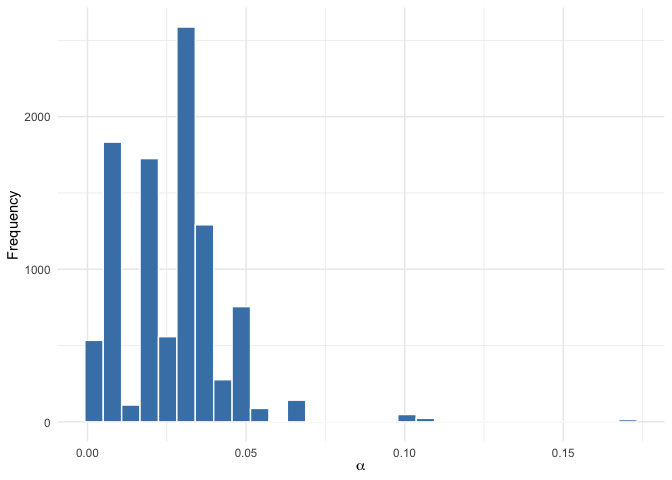
# Posterior distribution of rho (ranking positions)
plot(mod, parameter = "rho", items = c(1, 2, 3))
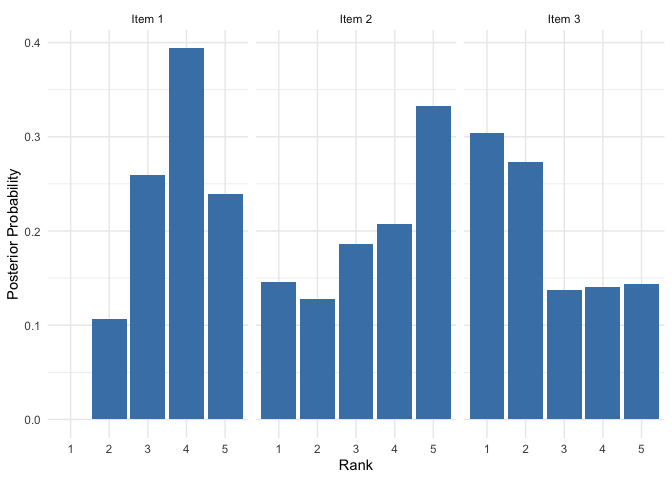
References
Sørensen, Øystein, Anja Stein, Waldir Leoncio Netto, and David S. Leslie. 2025. “Sequential Rank and Preference Learning with the Bayesian Mallows Model.” Bayesian Analysis, 1–26. https://doi.org/10.1214/25-BA1564.