Exploratory Chemometrics for Spectroscopy.
What is ChemoSpec?
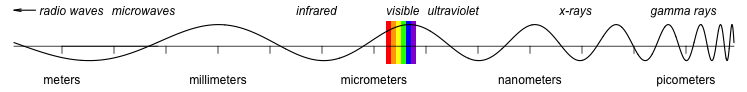
ChemoSpec is a collection of functions for top-down exploratory data analysis of spectral data including nuclear magnetic resonance (NMR), infrared (IR), Raman, X-ray fluorescence (XRF) and other similar types of spectroscopy. Includes functions for plotting and inspecting spectra, peak alignment, hierarchical cluster analysis (HCA), principal components analysis (PCA) and model-based clustering. Robust methods appropriate for this type of high-dimensional data are available. ChemoSpec is designed for structured experiments, such as metabolomics investigations, where the samples fall into treatment and control groups. Graphical output is formatted consistently for publication quality plots. ChemoSpec is intended to be very user friendly and to help you get usable results quickly. A vignette covering typical operations is available.
Learn more about ChemoSpec
Installing ChemoSpec from CRAN:
chooseCRANmirror() # choose a CRAN mirror
install.packages("ChemoSpec")
library("ChemoSpec")
Installing ChemoSpec from Github:
install.packages("remotes")
library("remotes")
install_github(repo = "bryanhanson/ChemoSpec@main")
library("ChemoSpec")
If you use @some_other_branch you can download other branches that might be available. They may or may not pass CRAN checks and thus may not install automatically using the method above. Check the NEWS file to see what's up.
Also...
ChemoSpec requires ChemoSpecUtils to work. It should install automatically, but if not, you can use a command similar to the above to install it.
To view the Vignettes:
To access the vignettes, use the following, or visit here.
browseVignettes("ChemoSpec")
Code of Conduct
This project is released with a Contributor Code of Conduct. By contributing, you agree to abide by its terms.
Contributing
If you would like to contribute to the project, please see Contributing Guide.
License Information
ChemoSpec is distributed under the GPL-3 license, as stated in the DESCRIPTION file. For more info, see the GPL site.
Questions? [email protected].
