A User-Friendly R 'shiny' Application for Multi-Omics Data Integration and Analysis.
Holomics

Holomics is an R Shiny application that enables users to perform single- and multi-omics analyses by providing a user-friendly interface to upload different omics datasets, select and run the implemented algorithms and finally visualize the generated results.
Holomics is primarily built on the R package mixOmics, which offers numerous algorithms for the integrative analysis of omics datasets. From this repertoire, the single-omics algorithms “Principal Component Analysis” (PCA) and “Partial Least Squares Discriminant Analysis” (PLS-DA), the pairwise-omics analysis “sparse Partial Least Squares” (sPLS) and the multi-omics framework DIABLO (“Data Integration Analysis for Biomarker discovery using Latent variable approaches for Omics studies”) have been implemented in Holomics.
Installation
CRAN
install.packages("Holomics")
Github
# Install devtools if it is not already installed
install.packages("devtools")
library(devtools)
# Install Holomics package
install_github("https://github.com/MolinLab/Holomics")
Additional packages
You need to install the Bioconductor package separately.
if (!require("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("mixOmics")
BiocManager::install("BiocParallel")
Start application
Either with
library(Holomics)
run_app()
or
Holomics::run_app()
Workflow
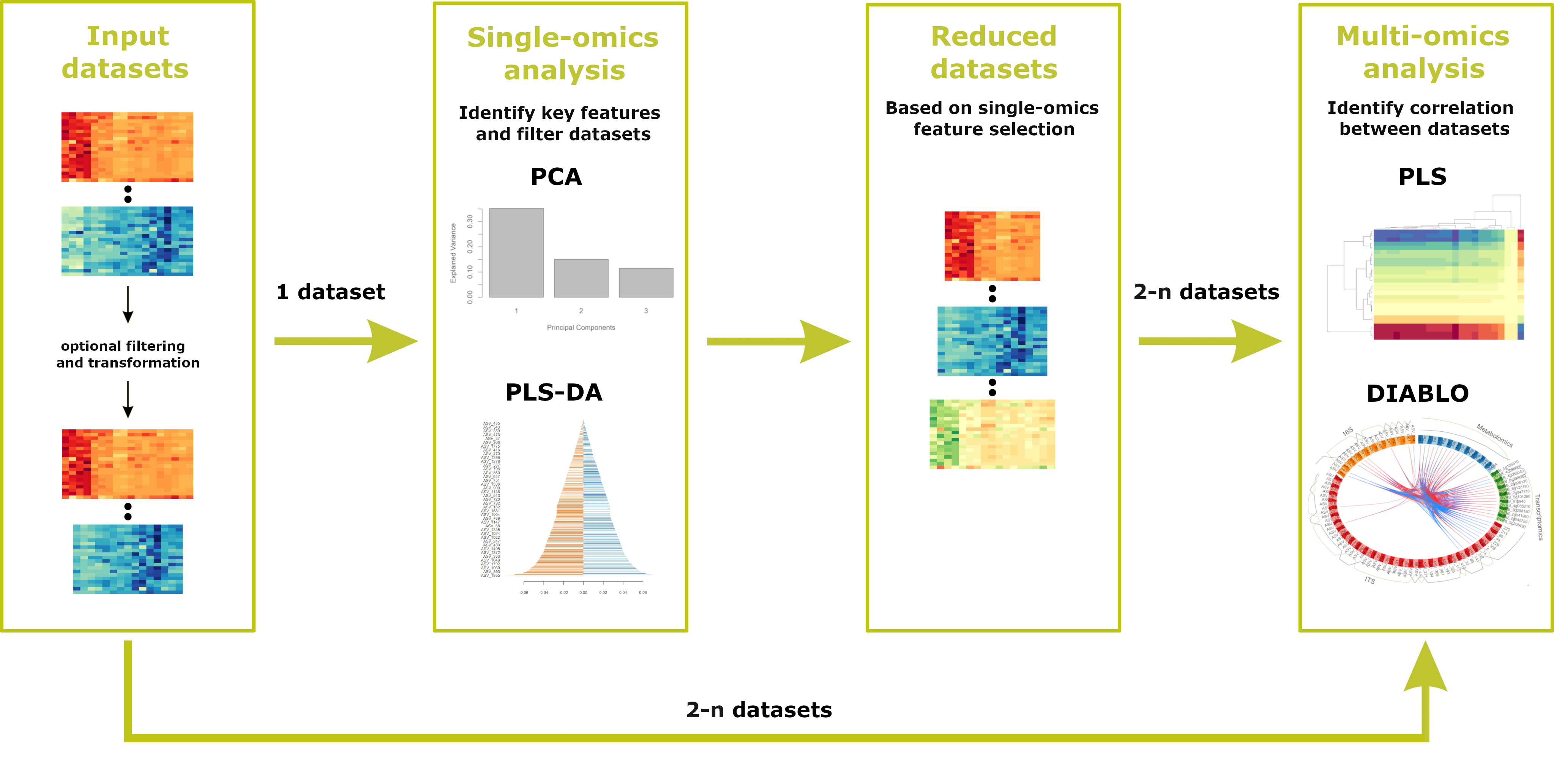
To use all the features offered, the following workflow should be followed. First, datasets are uploaded, during which any necessary pre-filtering or transformation steps take place. Next, the user should proceed to the single-omics analysis, where key features are identified and the datasets are reduced accordingly. After completing the single-omics analyses, the user can apply multi-omics analyses to identify correlations between two or more datasets. NOTE: If pre-filtered datasets (ideally generated earlier using Holomics) have already been uploaded, it is possible to start directly with the multi-omics analysis.
Further information
For further information on how to use Holomics please have a look at our vignette.


