EDGE Taxonomy Assignments Visualization.
MetaComp
Metagenome taxonomy assignment comparison toolkit. The toolkit is being developed for EDGE platform and reflects its backend specificity. The routines, however, can be used as a stand-alone library for multi-project comparative visualization of taxonomy assignments obtained for metagenomic samples processed with GOTTCHA/GOTTCHA2, BWA, KRAKEN, METAPHLAN, DIAMOND, or PANGIA. The heatmaps can be also visualized with this D3.js-based code which allows to see the exact abundance values in each cell.
0.0 Installation from CRAN
install.packages("MetaComp")
to use the library, simply load it into R environment:
library(MetaComp)
0.1 Installation from latest sources
install.packages("devtools")
library(devtools)
install_github(repo = 'seninp-bioinfo/MetaComp')
1.0 Reading a single taxonomic assignment files
the_gottcha2_assignment <- load_edge_assignment(data_file_g2, type = 'gottcha2')
the_kraken_assignment <- load_edge_assignment(data_file_k, type = 'kraken')
the_pangia_assignment <- load_edge_assignment(data_file_p, type = 'pangia')
1.1 Reading multiple taxonomic assignment files
The package functions load_xxx_assignments (where xxx stands for gottcha, kraken, or metaphlan) are designed to read a tool-specific assignment files. The configuration file for these functions must be tab-delimeted two columns file where the first column is the project id (used as the project's name in plotting), and the second column is an actual assignment file path:
the_assignments_list_g2 <- load_edge_assignments(config_file_g2, type = 'gottcha2')
the_assignments_list_k <- load_edge_assignments(config_file_k, type = 'kraken')
the_assignments_list_p <- load_edge_assignments(config_file_pangia, type = 'pangia')
2.0 Merging multiple taxonomic assignments into a single table
The merge_edge_assignments function is capable to merge a named list of GOTTCHA, Kraken, or MetaPhlAn assignments into a single table using LEVEL and TAXA columns as ids.
3.0 Plotting a single assignment as a heatmap
The function plot_edge_assignment accepts a single assignment table and outputs a ggplot object or produces a PDF plot using ggplot2's geom_tile.
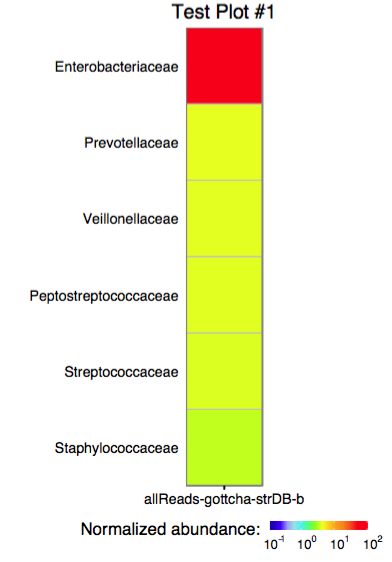
3.1 Plotting multiple assignments as a single heatmap
The function plot_merged_assignment accepts a single merged assignment table as an input and outputs a ggplot object or produces a PDF plot using ggplot2's geom_tile.
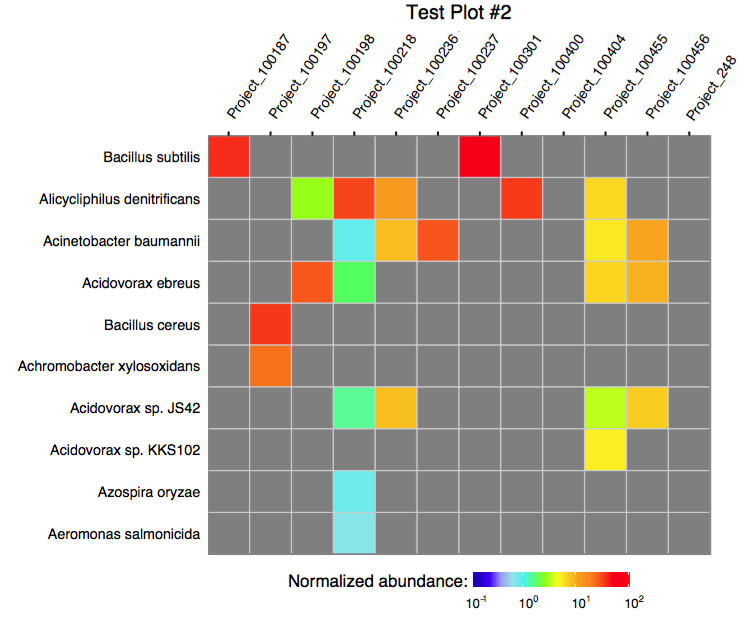
4.0. Running merge in a batch mode
The following script can be used to run the merge procedure in a batch mode:
# load library
require(MetaComp)
#
# configure runtime
options(echo = TRUE)
args <- commandArgs(trailingOnly = TRUE)
#
# print provided args
print(paste("provided args: ", args))
#
# acquire values
srcFile <- args[1]
destFile <- args[2]
taxonomyLevelArg <- args[3]
plotTitleArg <- args[4]
plotFileArg <- args[5]
#
# extended functionality was added in the release #3, and we don't want to break the legacy systems
#
if (length(args) > 5) {
rowLimitArg <- args[6]
sortingOrderArg <- args[7]
} else {
rowLimitArg <- 60
sortingOrderArg <- "abundance"
}
#
# read the data and produce the merged table
merged <- merge_edge_assignments(load_edge_assignments(srcFile, type = "gottcha2"))
#
# write the merge table as a TAB-delimeted file
write.table(merged, file = destFile, col.names = T, row.names = F, quote = T, sep = "\t")
#
# produce a PDF of the merged assignment
plot_merged_assignment(assignment = merged, taxonomy_level = taxonomyLevelArg,
sorting_order = sortingOrderArg, row_limit = base::strtoi(rowLimitArg),
plot_title = plotTitleArg, filename = plotFileArg)
To execute the scrip, use Rscript as shown below:
$> Rscript merge_and_plot_gottcha_assignments.R assignments_table_gottcha.txt merged_assignments.txt \
family "Merge test plot" merge_test 20 alphabetical
this command line arguments are (some of these are clickable -- so you can see examples):
Rscript- a way to execute the R scriptmerge_and_plot_gottcha_assignments.R- the above script filenameassignments_table_gottcha.txt- the tab delimeted table of assignments (two columns:project_idTABassignment_path)merged_assignments_gottcha.txt- the tab-delimeted output file namefamily- a LEVEL at which the plot should be produced"Merge test plot"- the output plot's titlemerge_test- the output plot filename mask,".pdf"and".svg"files will be produced...20the max number of rows to plot (in the specified sorting order)alphabeticalthe merged plot sorting order.
