Network Analysis for Omics Data.
MetaNet 
MetaNet: Network analysis for multi-omics
The HTML documentation of the latest version is available at Github page.
Tutorial📖
Please go to https://bookdown.org/Asa12138/metanet_book/ for the full vignette.

Installation
You can install the released version of MetaNet from CRAN with:
install.packages("MetaNet")
You can install the development version of MetaNet from GitHub with:
# install.packages("devtools")
devtools::install_github("Asa12138/MetaNet")
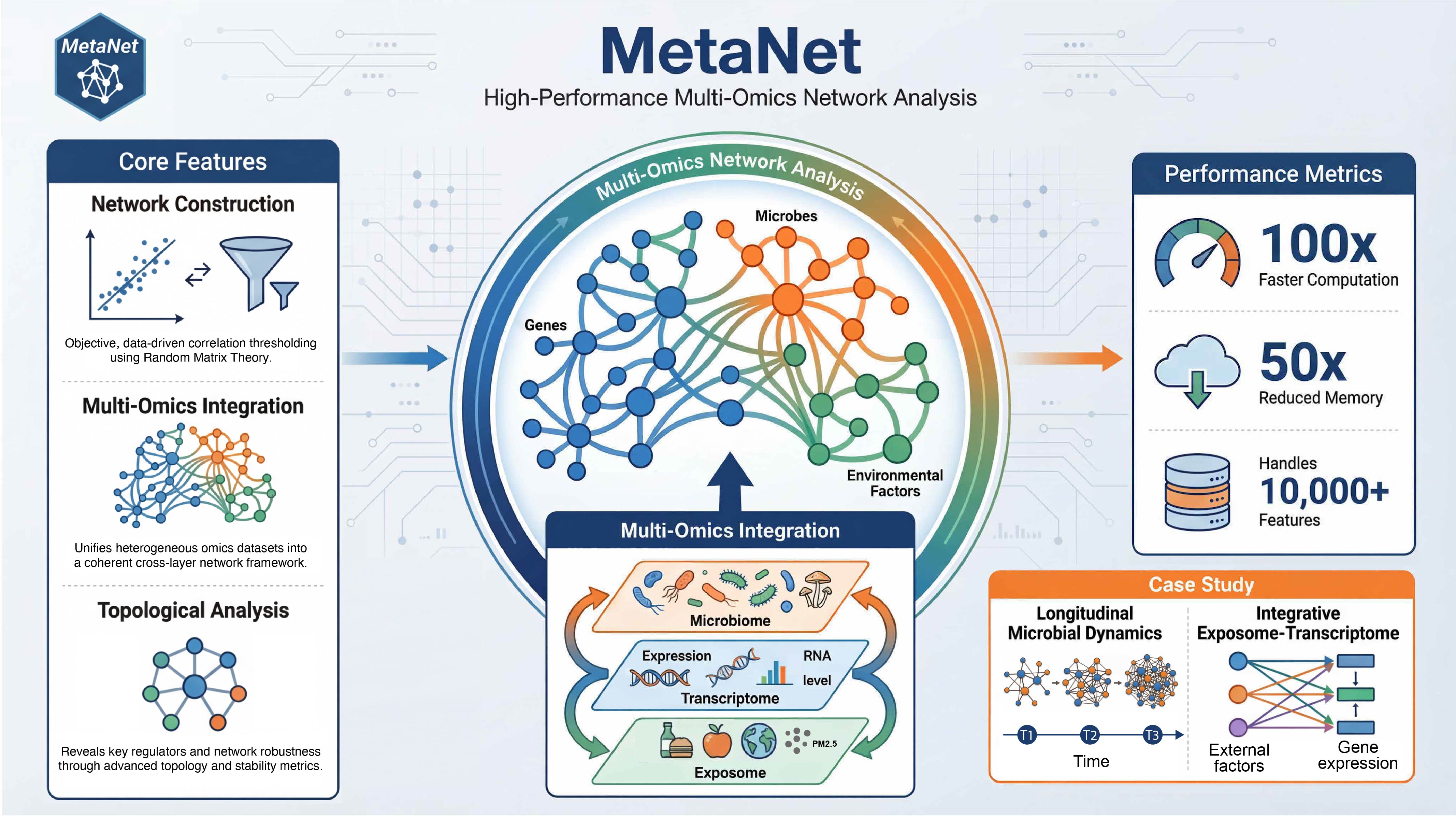
Workflow overview
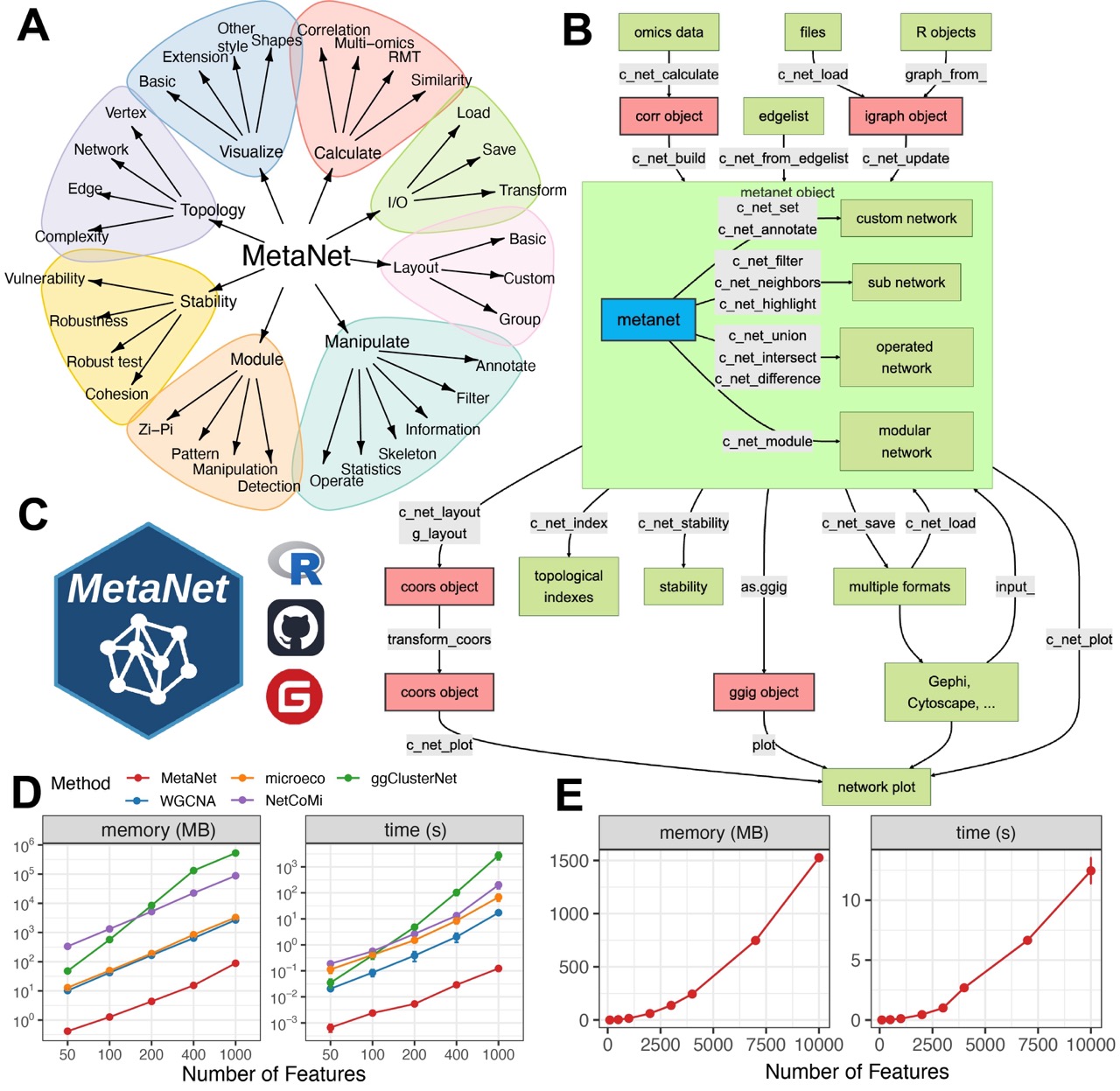
Figure 1. Overview of the MetaNet workflow and its high-efficiency computation.
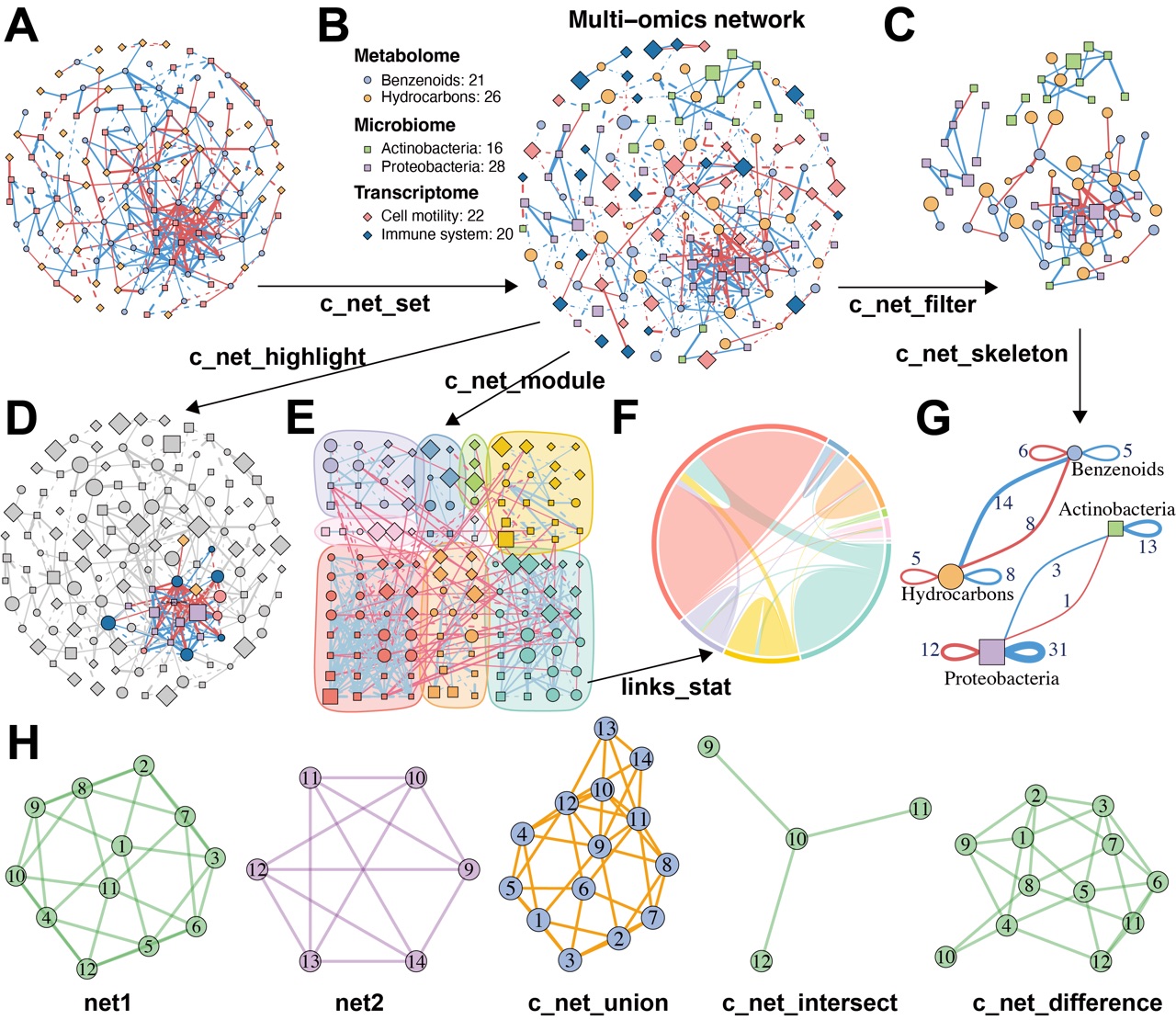
Figure 2. MetaNet supports flexible and intuitive network manipulation.
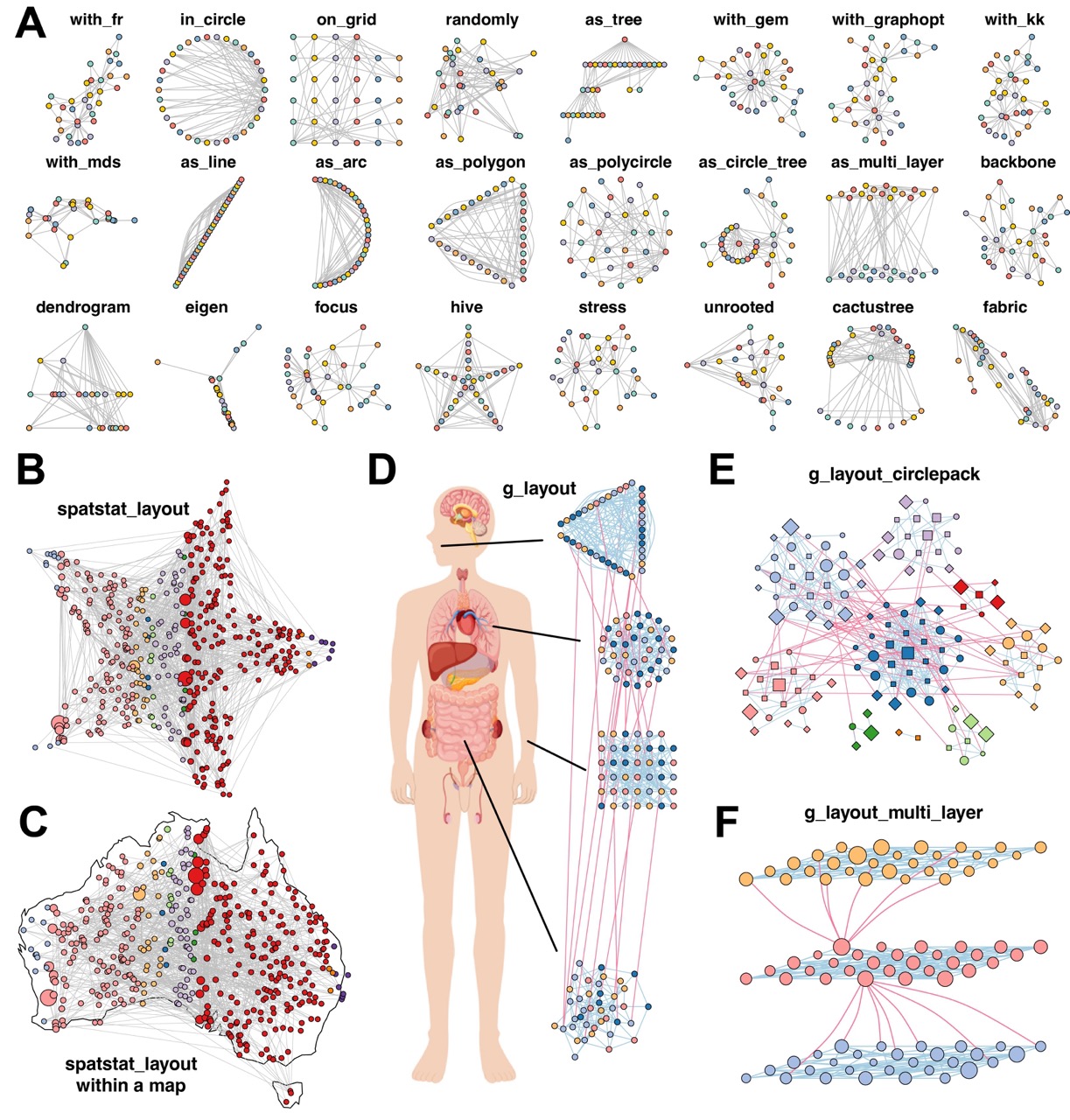
Figure 3. MetaNet enables diverse and powerful network layout strategies.
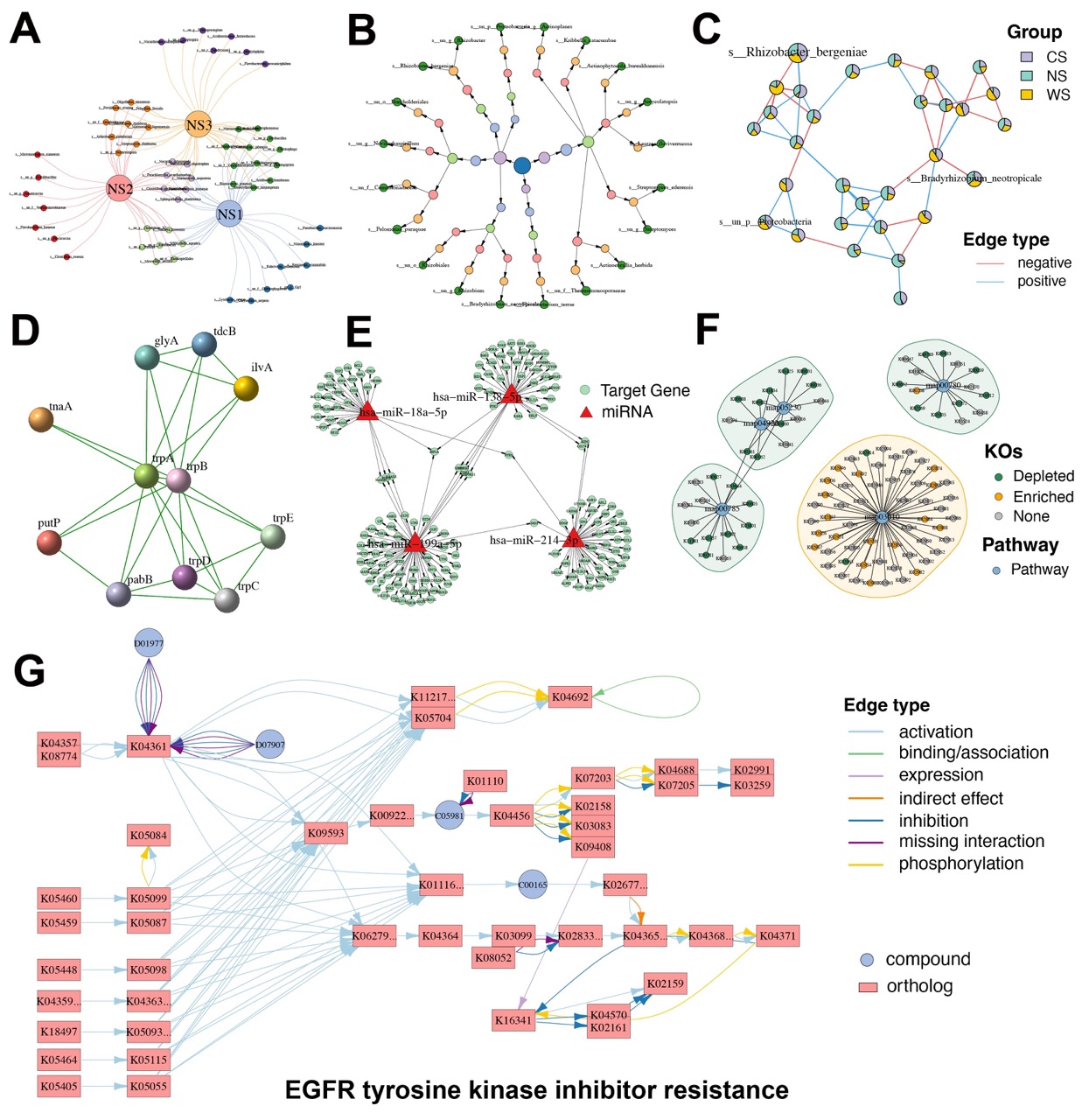
Figure 4. Diverse specialized network visualizations by MetaNet.
Citation
Please cite:
- Peng, C. et al. MetaNet: a scalable and integrated tool for reproducible omics network analysis. 2025.06.26.661636 Preprint at https://doi.org/10.1101/2025.06.26.661636 (2025).
Need helps?
If you have questions/issues, please visit MetaNet homepage first. Your problems are mostly documented. If you think you found a bug, please post on github issue.


