Processing and Physical Activity Summaries of Minute Level Activity Data.
arctools
- Installation
- Documentation
- Using
arctoolspackage to compute physical activity summaries - Additional
activity_statsmethod options - Components of
activity_statsmethod- <a href="#expand-the-length-of-minute-level-ac-vector-to-full-24-hour-periods-with-midnight_to_midnight" id="toc-expand-the-length-of-minute-level-ac-vector-to-full-24-hour-periods-with-midnight_to_midnight">Expand the length of minute-level AC vector to full 24-hour periods with
midnight_to_midnight - Get wear/non-wear flag with
get_wear_flag - Get valid/non-valid day flag with
get_valid_day_flag - Impute missing data with
impute_missing_data - Create PA characteristics with
summarize_PA
- <a href="#expand-the-length-of-minute-level-ac-vector-to-full-24-hour-periods-with-midnight_to_midnight" id="toc-expand-the-length-of-minute-level-ac-vector-to-full-24-hour-periods-with-midnight_to_midnight">Expand the length of minute-level AC vector to full 24-hour periods with
- Citation
The arctools package allows to generate summaries of the minute-level physical activity (PA) data. The default parameters are chosen for the Actigraph activity counts collected with a wrist-worn device; however, the package can be used for other minute-level PA data with the corresponding timepstamps vector.
Below, we demonstrate the use of arctools with the attached, exemplary minute-level Actigraph PA counts data.
Installation
You can install the released version of arctools from GitHub. Note you may need to install devtools package if not yet installed (the line commented below).
# install.packages("devtools")
devtools::install_github("martakarass/arctools")
Documentation
A PDF with detailed documentation of all methods can be accessed here.
Using arctools package to compute physical activity summaries
Reading PA data
Four CSV data sets with minute-level activity counts data are attached to the arctools package. The data file names are stored in extdata_fnames object that becomes available once the arctools package is loaded.
library(arctools)
library(data.table)
library(dplyr)
library(lubridate)
library(ggplot2)
## Read one of the data sets
fpath <- system.file("extdata", extdata_fnames[1], package = "arctools")
dat <- as.data.frame(fread(fpath))
rbind(head(dat, 3), tail(dat, 3))
#> Axis1 Axis2 Axis3 vectormagnitude timestamp
#> 1 1021 1353 2170 2754 2018-07-13 10:00:00
#> 2 1656 1190 2212 3009 2018-07-13 10:01:00
#> 3 2540 1461 1957 3524 2018-07-13 10:02:00
#> 10078 0 0 0 0 2018-07-20 09:57:00
#> 10079 0 0 0 0 2018-07-20 09:58:00
#> 10080 0 0 0 0 2018-07-20 09:59:00
The data columns are:
Axis1- sensor’s X axis minute-level counts data,Axis2- sensor’s Y axis minute-level counts data,Axis3- sensor’s Z axis minute-level counts data,vectormagnitude- minute-level counts data defined assqrt(Axis1^2 + Axis2^2 + Axis3^2),timestamp- time-stamps corresponding to minute-level measures.
## Plot activity counts
## Format timestamp data column from character to POSIXct object
ggplot(dat, aes(x = ymd_hms(timestamp), y = vectormagnitude)) +
geom_line(size = 0.3, alpha = 0.8) +
labs(x = "Time", y = "Activity counts") +
theme_gray(base_size = 10) +
scale_x_datetime(date_breaks = "1 day", date_labels = "%b %d")
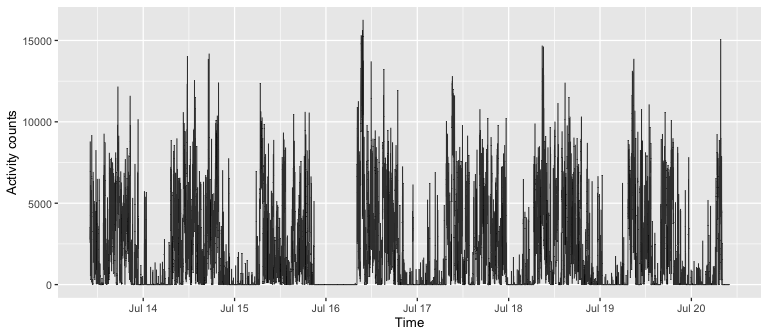
Computing summaries with activity_stats method
acc <- dat$vectormagnitude
acc_ts <- ymd_hms(dat$timestamp)
activity_stats(acc, acc_ts)
#> n_days n_valid_days wear_time_on_valid_days tac tlac ltac
#> 1 8 4 1440 2826648 6429.838 14.8546
#> astp satp time_spent_active time_spent_nonactive
#> 1 0.1781782 0.09516215 499.5 940.5
#> no_of_active_bouts no_of_nonactive_bouts mean_active_bout mean_nonactive_bout
#> 1 89 89.5 5.61236 10.50838
Output explained
To explain activity_stats method output, we first define the terms activity count, active/non-active minute, active/non-active bout, and valid day.
- Activity count (AC) - a minute-level metric of PA volume.
- Active minute - a minute with AC equal or above a fixed threshold; for wrist-worn Actigraph
we use AC>=1853 (method’s default). - Non-active (sedentary) minute - a minute with AC below a fixed threshold; for wrist-worn Actigraph
we use AC<1853 (method’s default). - Active bout - a sequence of 1 or more consecutive active minute(s).
- Non-active bout - a sequence of 1 or more consecutive non-active minute(s).
- Valid day - a day with no more than 10% of the non-wear time (see Details in
?activity_stats).
Meta information:
n_days- number of days (unique day dates) of data collection.n_valid_days- number of days (unique day dates) of data collection determined as valid days.wear_time_on_valid_days- average number of wear-time minutes across valid days.
Summaries of PA volumes metrics:
tac- TAC, Total activity counts per day - sum of AC measured on valid days divided by the number of valid days.tlac- TLAC, Total-log activity counts per day - sum of log(1+AC) measured on valid days divided by the number of valid days. Here ‘log’ denotes the natural logarithm.ltac- LTAC, Log-total activity counts - natural logarithm of TAC.time_spent_active- Average number of active minutes per valid day.time_spent_nonactive- Average number of sedentary minutes per valid day.
Summaries of PA fragmentation metrics:
astp- ASTP, active to sedentary transition probability on valid days.satp- SATP, sedentary to active transition probability on valid days.no_of_active_bouts- Average number of active minutes per valid day.no_of_nonactive_bouts- Average number of sedentary minutes per valid day.mean_active_bout- Average duration (in minutes) of an active bout on valid days.mean_nonactive_bout- Average duration (in minutes) of a sedentary bout on valid days.
Additionalactivity_stats method options
Summarizing PA within a fixed set of minutes only
The subset_minutes argument allows to specify a subset of a day’s minutes where activity summaries should be computed. There are 1440 minutes in a 24-hour day where 1 denotes 1st minute of the day (from 00:00 to 00:01), and 1440 denotes the last minute (from 23:59 to 00:00).
Here, we summarize PA observed between 12:00 AM and 6:00 AM.
subset_12am_6am <- 1 : (6 * 1440/24)
activity_stats(acc, acc_ts, subset_minutes = subset_12am_6am)
#> n_days n_valid_days wear_time_on_valid_days tac_0to6only tlac_0to6only
#> 1 8 4 1440 65477.5 322.1523
#> ltac_0to6only astp_0to6only satp_0to6only time_spent_active_0to6only
#> 1 11.08946 0.5581395 0.02004295 10.75
#> time_spent_nonactive_0to6only no_of_active_bouts_0to6only
#> 1 349.25 6
#> no_of_nonactive_bouts_0to6only mean_active_bout_0to6only
#> 1 7 1.791667
#> mean_nonactive_bout_0to6only
#> 1 49.89286
By default, column names have a suffix added to denote that a subset of minutes was used (here, _0to6only). This can be disabled by setting adjust_out_colnames to FALSE.
subset_12am_6am = 1 : (6/24 * 1440)
subset_6am_12pm = (6/24 * 1440 + 1) : (12/24 * 1440)
subset_12pm_6pm = (12/24 * 1440 + 1) : (18/24 * 1440)
subset_6pm_12am = (18/24 * 1440 + 1) : (24/24 * 1440)
out <- rbind(
activity_stats(acc, acc_ts, subset_minutes = subset_12am_6am, adjust_out_colnames = FALSE),
activity_stats(acc, acc_ts, subset_minutes = subset_6am_12pm, adjust_out_colnames = FALSE),
activity_stats(acc, acc_ts, subset_minutes = subset_12pm_6pm, adjust_out_colnames = FALSE),
activity_stats(acc, acc_ts, subset_minutes = subset_6pm_12am, adjust_out_colnames = FALSE))
rownames(out) <- c("12am-6am", "6am-12pm", "12pm-6pm", "6pm-12am")
out
#> n_days n_valid_days wear_time_on_valid_days tac tlac
#> 12am-6am 8 4 1440 65477.5 322.1523
#> 6am-12pm 8 4 1440 1089788.5 2139.4534
#> 12pm-6pm 8 4 1440 994104.8 2194.8539
#> 6pm-12am 8 4 1440 677277.5 1773.3781
#> ltac astp satp time_spent_active time_spent_nonactive
#> 12am-6am 11.08946 0.5581395 0.02004295 10.75 349.25
#> 6am-12pm 13.90149 0.1501377 0.15406162 181.50 178.50
#> 12pm-6pm 13.80960 0.1751337 0.18641618 187.00 173.00
#> 6pm-12am 13.42584 0.2037422 0.10323253 120.25 239.75
#> no_of_active_bouts no_of_nonactive_bouts mean_active_bout
#> 12am-6am 6.00 7.00 1.791667
#> 6am-12pm 27.25 27.50 6.660550
#> 12pm-6pm 32.75 32.25 5.709924
#> 6pm-12am 24.50 24.75 4.908163
#> mean_nonactive_bout
#> 12am-6am 49.892857
#> 6am-12pm 6.490909
#> 12pm-6pm 5.364341
#> 6pm-12am 9.686869
Summarizing PA within a subset of weekdays only
The subset_weekdays argument allows to specify days of a week within which activity summaries are to be computed; it takes values between 1 (Sunday) to 7 (Saturday). Default is NULL (all days of a week are used).
Here, we summarize PA within weekday days only. Note that in the method output, then_daysandn_valid_dayscolumns only count the days from the selected week days subset; for example, below, n_days number of unique day dates in data is 6 despite the range of data collection without subsetting ranges 8 days.
# day of a week indices 2,3,4,5,6 correspond to Mon,Tue,Wed,Thu,Fri
subset_weekdays <- c(2:6)
activity_stats(acc, acc_ts, subset_weekdays = subset_weekdays)
#> n_days n_valid_days wear_time_on_valid_days tac_weekdays23456only
#> 1 6 3 1440 2865711
#> tlac_weekdays23456only ltac_weekdays23456only astp_weekdays23456only
#> 1 6444.155 14.86833 0.1757294
#> satp_weekdays23456only time_spent_active_weekdays23456only
#> 1 0.09459459 502.6667
#> time_spent_nonactive_weekdays23456only no_of_active_bouts_weekdays23456only
#> 1 937.3333 88.33333
#> no_of_nonactive_bouts_weekdays23456only mean_active_bout_weekdays23456only
#> 1 88.66667 5.690566
#> mean_nonactive_bout_weekdays23456only
#> 1 10.57143
Note the subset_weekdays argument can be combined with other arguments, i.e. subset_minutes to subset of a day’s minutes where activity summaries should be computed.
# day of a week indices 7,1 correspond to Sat,Sun
subset_weekdays <- c(7,1)
activity_stats(acc, acc_ts, subset_weekdays = subset_weekdays, subset_minutes = subset_6am_12pm)
#> n_days n_valid_days wear_time_on_valid_days tac_6to12only_weekdays17only
#> 1 2 1 1440 917368
#> tlac_6to12only_weekdays17only ltac_6to12only_weekdays17only
#> 1 2071.864 13.72926
#> astp_6to12only_weekdays17only satp_6to12only_weekdays17only
#> 1 0.1840491 0.1522843
#> time_spent_active_6to12only_weekdays17only
#> 1 163
#> time_spent_nonactive_6to12only_weekdays17only
#> 1 197
#> no_of_active_bouts_6to12only_weekdays17only
#> 1 30
#> no_of_nonactive_bouts_6to12only_weekdays17only
#> 1 30
#> mean_active_bout_6to12only_weekdays17only
#> 1 5.433333
#> mean_nonactive_bout_6to12only_weekdays17only
#> 1 6.566667
Summarizing PA with a fixed set of minutes excluded
The exclude_minutes argument allows specifying a subset of a day’s minutes excluded for computing activity summaries.
Here, we summarize PA while excluding observations between 11:00 PM and 5:00 AM.
subset_11pm_5am <- c(
(23 * 1440/24 + 1) : 1440, ## 11:00 PM - midnight
1 : (5 * 1440/24) ## midnight - 5:00 AM
)
activity_stats(acc, acc_ts, exclude_minutes = subset_11pm_5am)
#> n_days n_valid_days wear_time_on_valid_days tac_23to5removed
#> 1 8 4 1440 2735749
#> tlac_23to5removed ltac_23to5removed astp_23to5removed satp_23to5removed
#> 1 6052.84 14.82192 0.1702018 0.1395057
#> time_spent_active_23to5removed time_spent_nonactive_23to5removed
#> 1 483.25 596.75
#> no_of_active_bouts_23to5removed no_of_nonactive_bouts_23to5removed
#> 1 82.25 83.25
#> mean_active_bout_23to5removed mean_nonactive_bout_23to5removed
#> 1 5.87538 7.168168
Summarizing PA with in-bed time excluded
The in_bed_time and out_bed_time arguments allow to provide day-specific in-bed periods to be excluded from analysis.
Here, we summarize PA excluding in-bed time estimated by ActiLife software.
ActiLife-estimated in-bed data
The ActiLife-estimated in-bed data file is attached to the arctools package. The sleep data columns include:
Subject Name- subject IDs corresponding to AC data, stored inextdata_fnames,In Bed Time- ActiLife-estimated start of in-bed interval for each day of the measurement,Out Bed Time- ActiLife-estimated end of in-bed interval.
## Read sleep details data file
SleepDetails_fname <- "BatchSleepExportDetails_2020-05-01_14-00-46.csv"
SleepDetails_fpath <- system.file("extdata", SleepDetails_fname, package = "arctools")
SleepDetails <- as.data.frame(fread(SleepDetails_fpath))
## Filter sleep details data to keep ID1 file
SleepDetails_sub <-
SleepDetails %>%
filter(`Subject Name` == "ID_1") %>%
select(`Subject Name`, `In Bed Time`, `Out Bed Time`)
str(SleepDetails_sub)
#> 'data.frame': 6 obs. of 3 variables:
#> $ Subject Name: chr "ID_1" "ID_1" "ID_1" "ID_1" ...
#> $ In Bed Time : chr "7/13/2018 9:18:00 PM" "7/14/2018 10:41:00 PM" "7/16/2018 7:46:00 PM" "7/17/2018 11:30:00 PM" ...
#> $ Out Bed Time: chr "7/14/2018 4:50:00 AM" "7/15/2018 5:40:00 AM" "7/17/2018 4:32:00 AM" "7/18/2018 6:32:00 AM" ...
We transform dates stored as character into POSIXct object, and then use in/out-bed dates vectors in activity_stats method.
in_bed_time <- mdy_hms(SleepDetails_sub[, "In Bed Time"])
out_bed_time <- mdy_hms(SleepDetails_sub[, "Out Bed Time"])
activity_stats(acc, acc_ts, in_bed_time = in_bed_time, out_bed_time = out_bed_time)
#> n_days n_valid_days wear_time_on_valid_days tac_inbedremoved
#> 1 8 4 1440 2746582
#> tlac_inbedremoved ltac_inbedremoved astp_inbedremoved satp_inbedremoved
#> 1 6062.753 14.82587 0.1703551 0.1580934
#> time_spent_active_inbedremoved time_spent_nonactive_inbedremoved
#> 1 485.75 529.75
#> no_of_active_bouts_inbedremoved no_of_nonactive_bouts_inbedremoved
#> 1 82.75 83.75
#> mean_active_bout_inbedremoved mean_nonactive_bout_inbedremoved
#> 1 5.870091 6.325373
Components of activity_stats method
The primary method activity_stats is composed of several steps implemented in their respective functions. Below, we demonstrate how to produce activity_stats results step by step with these functions.
We reuse the objects:
acc- a numeric vector; minute-level activity counts data,acc_ts- aPOSIXctvector; minute-level time ofaccdata collection.
df <- data.frame(acc = acc, acc_ts = acc_ts)
rbind(head(df, 3), tail(df, 3))
#> acc acc_ts
#> 1 2754 2018-07-13 10:00:00
#> 2 3009 2018-07-13 10:01:00
#> 3 3524 2018-07-13 10:02:00
#> 10078 0 2018-07-20 09:57:00
#> 10079 0 2018-07-20 09:58:00
#> 10080 0 2018-07-20 09:59:00
Expand the length of minute-level AC vector to full 24-hour periods with midnight_to_midnight
- In the returned vector, the first observation corresponds to the minute of
00:00-00:01on the first day of data collection, and the last observation corresponds to the minute of23:50-00:00on the last day of data collection. - Entries corresponding to non-measured minutes are filled with
NA.
Here, collected data cover total of 7*24*1440 = 10080 minutes (from 2018-07-13 10:00:00 to 2018-07-20 09:59:00), but spans 8*24*1440 = 11520 minutes of full midnight-to-midnight days (from 2018-07-13 00:00:00 to 2018-07-20 23:59:00).
acc <- midnight_to_midnight(acc = acc, acc_ts = acc_ts)
## Vector length on non NA-obs, vector length after acc
c(length(acc[!is.na(acc)]), length(acc))
#> [1] 10080 11520
Get wear/non-wear flag with get_wear_flag
Function get_wear_flag computes wear/non-wear flag (1/0) for each minute of activity counts data. Method implements wear/non-wear detection algorithm closely following that of Choi et al. (2011). See ?get_wear_flag for more details and function arguments.
- The returned vector has value
1for wear and0for non-wear flagged minute. - If there is an
NAentry in a data input vector, then the returned vector will have a corresponding entry set toNAtoo.
wear_flag <- get_wear_flag(acc)
## Proportion of wear time across the days
wear_flag_mat <- matrix(wear_flag, ncol = 1440, byrow = TRUE)
round(apply(wear_flag_mat, 1, sum, na.rm = TRUE) / 1440, 3)
#> [1] 0.583 1.000 0.874 0.679 1.000 1.000 1.000 0.338
Get valid/non-valid day flag with get_valid_day_flag
Function get_valid_day_flag computes valid/non-valid day flag (1/0) for each minute of activity counts data. See ?get_valid_day_flag for more details and function arguments.
Here, 4 out of 8 days have more than 10% (144 minutes) of missing data.
valid_day_flag <- get_valid_day_flag(wear_flag)
## Compute number of valid days
valid_day_flag_mat <- matrix(valid_day_flag, ncol = 1440, byrow = TRUE)
apply(valid_day_flag_mat, 1, mean, na.rm = TRUE)
#> [1] 0 1 0 0 1 1 1 0
Impute missing data with impute_missing_data
Function impute_missing_data imputes missing data in valid days based on the “average day profile”, a minute-wise average of wear-time AC across valid days. See ?get_valid_day_flag for more details and function arguments.
## Copies of original objects for the purpose of demonstration
acc_cpy <- acc
wear_flag_cpy <- wear_flag
## Artificially replace 1h (4%) of a valid day with non-wear
repl_idx <- seq(from = 1441, by = 1, length.out = 60)
acc_cpy[repl_idx] <- 0
wear_flag_cpy[repl_idx] <- 0
## Impute data for minutes identified as non-wear in days identified as valid
acc_cpy_imputed <- impute_missing_data(acc_cpy, wear_flag_cpy, valid_day_flag)
## Compare mean activity count on valid days before and after imputation
c(mean(acc_cpy[which(valid_day_flag == 1)]),
mean(acc_cpy_imputed[which(valid_day_flag == 1)]))
#> [1] 1955.521 1957.186
Create PA characteristics with summarize_PA
Finally, method summarize_PA computes PA summaries. Similarly as activity_stats, it accepts arguments to subset/exclude minutes. See ?activity_stats for more details and function arguments.
summarize_PA(acc, acc_ts, wear_flag, valid_day_flag)
#> n_days n_valid_days wear_time_on_valid_days tac tlac ltac
#> 1 8 4 1440 2826648 6429.838 14.8546
#> astp satp time_spent_active time_spent_nonactive
#> 1 0.1781782 0.09516215 499.5 940.5
#> no_of_active_bouts no_of_nonactive_bouts mean_active_bout mean_nonactive_bout
#> 1 89 89.5 5.61236 10.50838
It returns the same results as the activity_stats function:
activity_stats(dat$vectormagnitude, ymd_hms(dat$timestamp))
#> n_days n_valid_days wear_time_on_valid_days tac tlac ltac
#> 1 8 4 1440 2826648 6429.838 14.8546
#> astp satp time_spent_active time_spent_nonactive
#> 1 0.1781782 0.09516215 499.5 940.5
#> no_of_active_bouts no_of_nonactive_bouts mean_active_bout mean_nonactive_bout
#> 1 89 89.5 5.61236 10.50838
Citation
citation("arctools")
#>
#> To cite arctools in publications use:
#>
#> Karas, M., Schrack, J., and Urbanek, J. (2021). arctools: Processing
#> and Physical Activity Summaries of Minute Level Activity Data. R
#> package version 1.1.4.
#>
#> A BibTeX entry for LaTeX users is
#>
#> @Manual{,
#> title = {{arctools: Processing and Physical Activity Summaries of Minute Level Activity Data}},
#> author = {Marta Karas and Jennifer Schrack and Jacek Urbanek},
#> url = {https://CRAN.R-project.org/package=arctools},
#> note = {R package version 1.1.4},
#> year = {2021},
#> }