Description
Bayesian Estimation of Naloxone Kit Number Under-Reporting.
Description
Bayesian model and associated tools for generating estimates of total naloxone kit numbers distributed and used from naloxone kit orders data. Provides functions for generating simulated data of naloxone kit use and functions for generating samples from the posterior.
README.md
bennu 
Bayesian Estimation of Naloxone Numbers Underreporting (BENNU)
The package name comes from the Welsh word for “to finish” (pronounced benn-y)
An R package 📦 for generating estimates of total naloxone kit numbers distributed and used from naloxone kit orders data.
Installation
You can install the released version of bennu from CRAN with:
install.packages("bennu")
And the development version from GitHub with:
# install.packages("devtools")
devtools::install_github("sempwn/bennu")
Example
This example runs output for test data generated by the package:
library(bennu)
library(rstan)
#> Loading required package: StanHeaders
#>
#> rstan version 2.26.22 (Stan version 2.26.1)
#> For execution on a local, multicore CPU with excess RAM we recommend calling
#> options(mc.cores = parallel::detectCores()).
#> To avoid recompilation of unchanged Stan programs, we recommend calling
#> rstan_options(auto_write = TRUE)
#> For within-chain threading using `reduce_sum()` or `map_rect()` Stan functions,
#> change `threads_per_chain` option:
#> rstan_options(threads_per_chain = 1)
#> Do not specify '-march=native' in 'LOCAL_CPPFLAGS' or a Makevars file
library(bayesplot)
#> This is bayesplot version 1.10.0
#> - Online documentation and vignettes at mc-stan.org/bayesplot
#> - bayesplot theme set to bayesplot::theme_default()
#> * Does _not_ affect other ggplot2 plots
#> * See ?bayesplot_theme_set for details on theme setting
rstan_options(auto_write = TRUE)
options(mc.cores = parallel::detectCores(logical = FALSE))
## basic example code
d <- generate_model_data()
# note iter should be at least 2000 to generate a reasonable posterior sample
fit <- est_naloxone(d,iter=500)
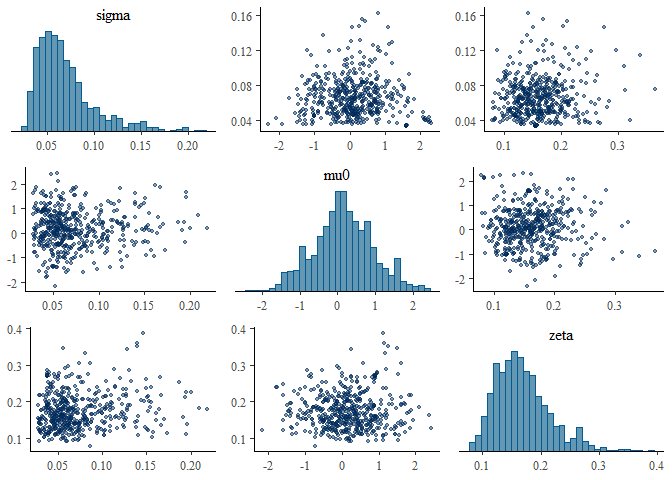
mcmc_pairs(fit, pars = c("sigma","mu0","zeta"),
off_diag_args = list(size = 1, alpha = 0.5))
Getting help
If you encounter a clear bug, please file an issue with a minimal reproducible example on GitHub.

