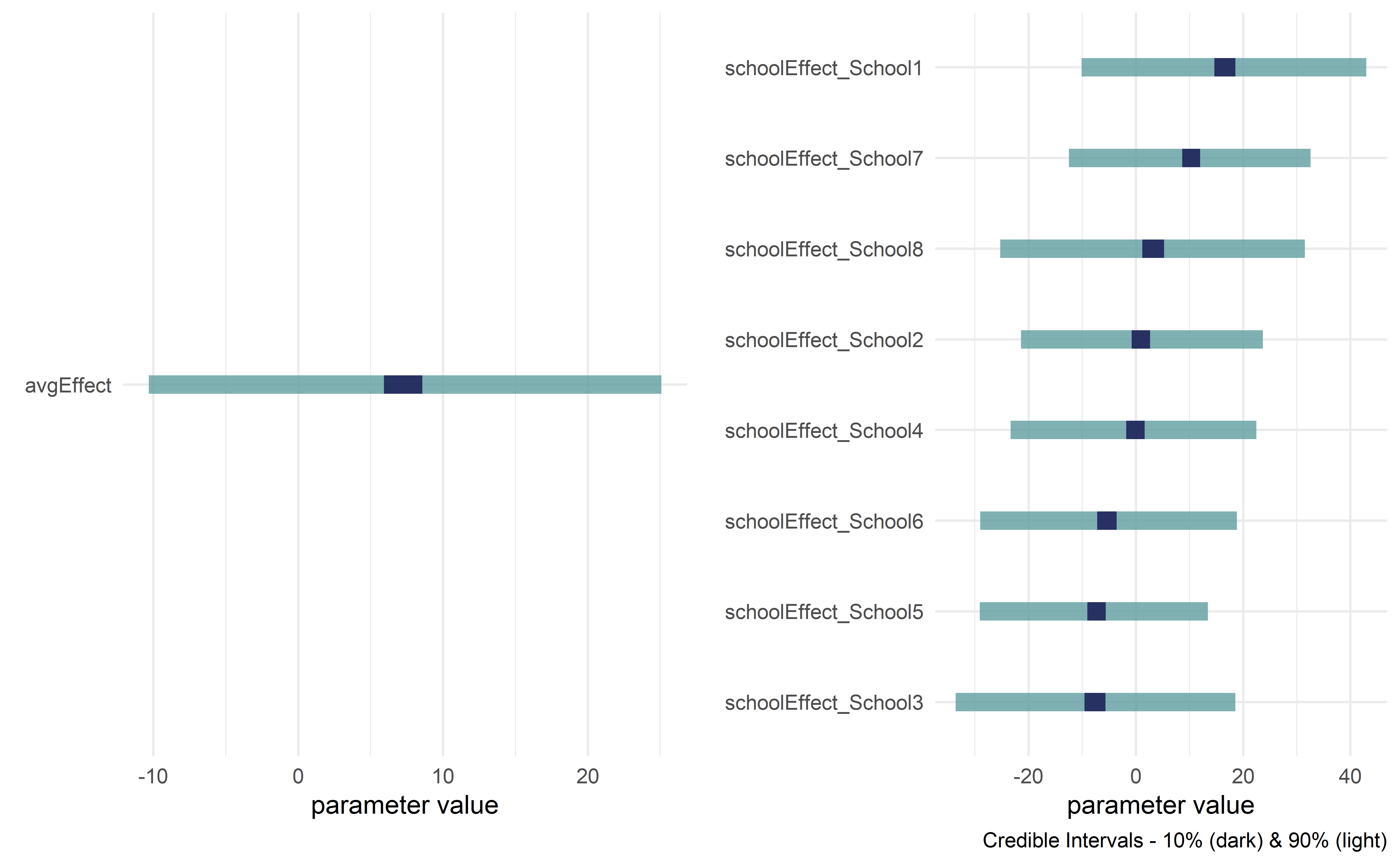Fast, Easy, and Visual Bayesian Inference.
causact
Accelerate computational Bayesian inference workflows in R through interactive modelling, visualization, and inference. The package leverages directed acyclic graphs (DAGs) to create a unified language language for business stakeholders, statisticians, and programmers. Due to its visual nature and simple model construction, causact serves as a great entry-point for newcomers to computational Bayesian inference.
The
causactpackage offers robust support for both foundational and advanced Bayesian models. While introductory models are well-covered, the utilization of multi-variate distributions such as multi-variate normal, multi-nomial, or dirichlet distributions, may not work as expected. There are ongoing enhancements in the pipeline to facilitate construction of these more intricate models.
While proficiency in R is the only requirement for users of this package, it also functions as a introductory probabilistic programming language, seamlessly incorporating the numpyro Python package to facilitate Bayesian inference without the need to learn any syntax outside of the package or R. Furthermore, the package streamlines the process of indexing categorical variables, which often presents a complex syntax hurdle for those new to computational Bayesian methods.
For enhanced learning, the causact package for Bayesian inference is featured in A Business Analyst's Introduction to Business Analytics available at https://www.causact.com/.
Feedback and encouragement is appreciated via github issues.
Installation Guide
To install the causact package, follow the steps outlined below:
CRAN Release Version (Recommended)
For the current stable release, which is tailored to integrate with Python’s numpyro package, employ the following command:
install.packages("causact")
Then, see Essential Dependencies if you want to be able to automate sampling using the numpyro package.
Development Release
If you want the most recent development version (not recommended), execute the following:
install.packages("remotes")
remotes::install_github("flyaflya/causact")
Essential Dependencies
To harness the full potential of causact for DAG visualization and Bayesian posterior analysis, it’s vital to ensure proper integration with the numpyro package. Given the Python-based nature of numpyro, a few essential dependencies must be in place. Execute the following commands after installing causact:
library(causact)
install_causact_deps()
If prompted, respond with Y to any inquiries related to installing miniconda.
Note: If opting for installation on Posit Cloud, temporarily adjust your project’s RAM to 4GB during the installation process (remember to APPLY CHANGES). This preemptive measure helps avoid encountering an
Error: Error creating conda environment [exit code 137]. After installation, feel free to revert the settings to 1GB of RAM.
Note: The September 11, 2023 release of
reticulate(v1.32) has caused an issue which gives aTypeError: the first argument must be callableerror when usingdag_numpyro()on windows. If you experience this, install the dev version ofreticulateby following the below steps:
Install RTOOLS by using installer at: https://cran.r-project.org/bin/windows/Rtools/
Run this to get the dev version of
reticulate:
# install DEV version of reticulate
# install.packages("pak") #uncomment as needed
pak::pak("rstudio/reticulate")
Retrograde Compatibility (Not Advised)
In cases where legacy compatibility is paramount and you still rely on the operationality of the dag_greta() function, consider installing v0.4.2 of the causact package. However, it’s essential to emphasize that this approach is not recommended for general usage:
### EXERCISE CAUTION BEFORE EXECUTING THESE LINES
### Only proceed if dag_greta() is integral to your existing codebase
install.packages("remotes")
remotes::install_github("flyaflya/[email protected]")
Your judicious choice of installation method will ensure a seamless and effective integration of the causact package into your computational toolkit.
Usage
Example taken from https://www.causact.com/graphical-models-tell-joint-distribution-stories.html#graphical-models-tell-joint-distribution-stories with the packages dag_foo() functions further described here:
Create beautiful model visualizations.
#> Initializing python, numpyro, and other dependencies. This may take up to 15 seconds...
#> Initializing python, numpyro, and other dependencies. This may take up to 15 seconds...COMPLETED!
#>
#> Attaching package: 'causact'
#> The following objects are masked from 'package:stats':
#>
#> binomial, poisson
#> The following objects are masked from 'package:base':
#>
#> beta, gamma
library(causact)
graph = dag_create() %>%
dag_node(descr = "Get Card", label = "y",
rhs = bernoulli(theta),
data = carModelDF$getCard) %>%
dag_node(descr = "Card Probability", label = "theta",
rhs = beta(2,2),
child = "y") %>%
dag_plate(descr = "Car Model", label = "x",
data = carModelDF$carModel,
nodeLabels = "theta",
addDataNode = TRUE)
graph %>% dag_render()
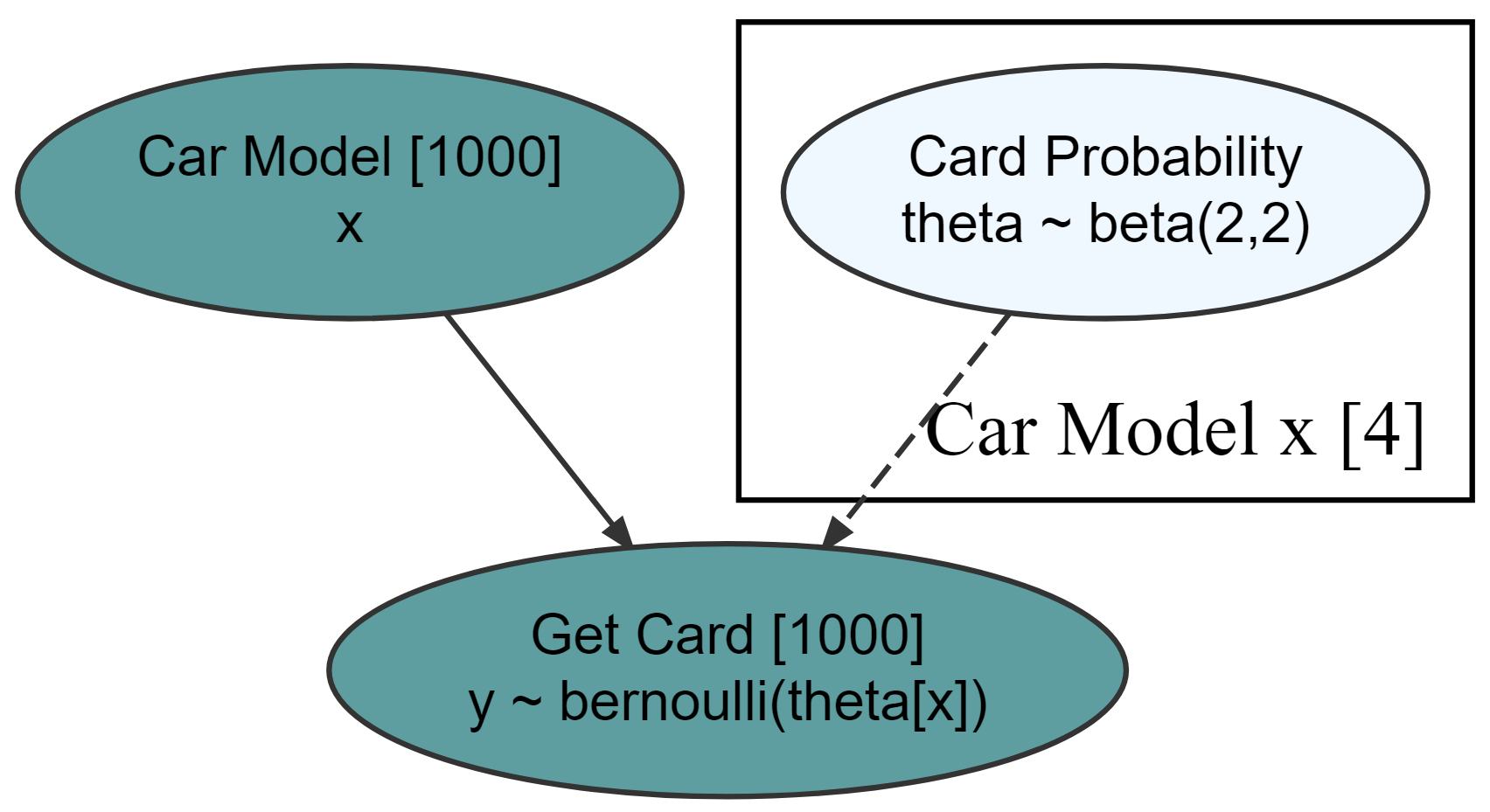
Hide model complexity, as appropriate, from domain experts and other less statistically minded stakeholders.
graph %>% dag_render(shortLabel = TRUE)
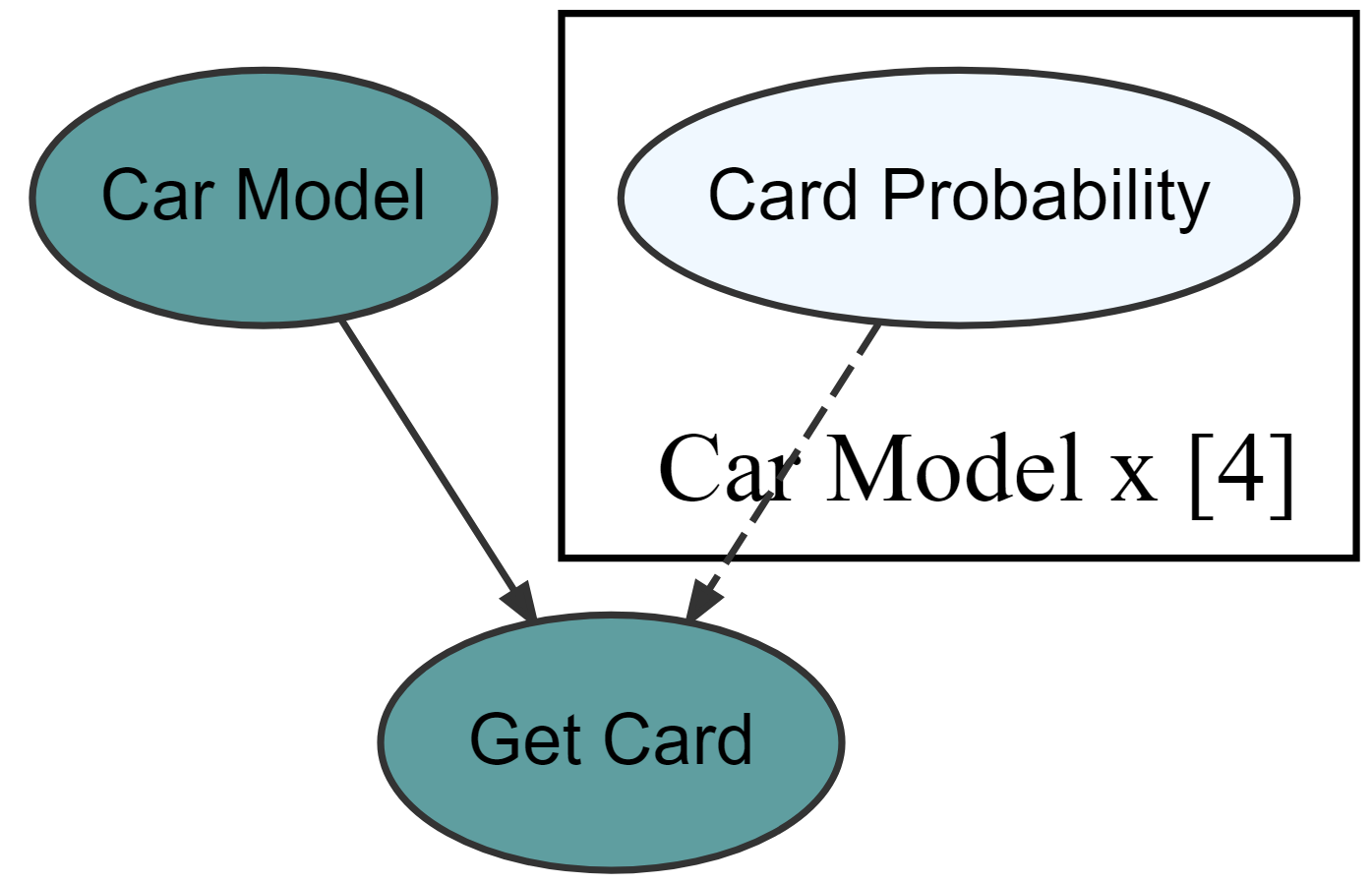
Get posterior while automatically running the underlying numpyro code
drawsDF = graph %>% dag_numpyro()
drawsDF ### see top of data frame
#> # A tibble: 4,000 × 4
#> theta_Toyota.Corolla theta_Subaru.Outback theta_Kia.Forte theta_Jeep.Wrangler
#> <dbl> <dbl> <dbl> <dbl>
#> 1 0.215 0.582 0.226 0.839
#> 2 0.194 0.643 0.264 0.853
#> 3 0.216 0.602 0.243 0.831
#> 4 0.217 0.647 0.177 0.843
#> 5 0.186 0.580 0.309 0.853
#> 6 0.230 0.633 0.237 0.846
#> 7 0.217 0.619 0.263 0.859
#> 8 0.205 0.692 0.274 0.862
#> 9 0.191 0.693 0.332 0.847
#> 10 0.188 0.655 0.174 0.851
#> # ℹ 3,990 more rows
Note: if you have used older versions of causact, please know that dag_numpyro() is a drop-in replacement for dag_greta().
Get quick view of posterior distribution
drawsDF %>% dagp_plot()
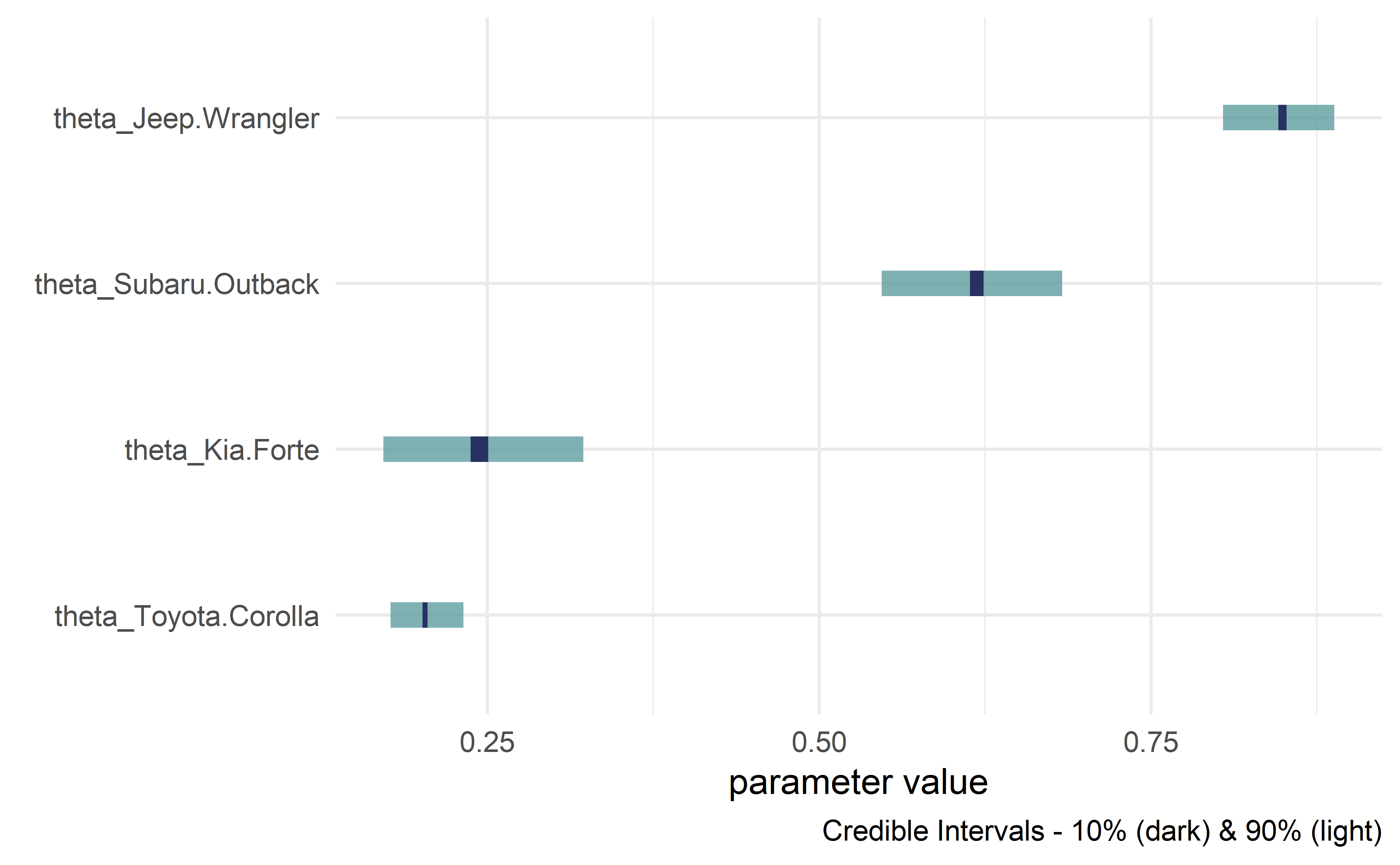
Credible interval plots.
OPTIONAL: See numpyro code without executing it (for debugging or learning)
numpyroCode = graph %>% dag_numpyro(mcmc = FALSE)
#>
#> ## The below code will return a posterior distribution
#> ## for the given DAG. Use dag_numpyro(mcmc=TRUE) to return a
#> ## data frame of the posterior distribution:
#> reticulate::py_run_string("
#> import numpy as np
#> import numpyro as npo
#> import numpyro.distributions as dist
#> import pandas as pd
#> from jax import random
#> from numpyro.infer import MCMC, NUTS
#> from jax.numpy import transpose as t
#> from jax.numpy import (exp, log, log1p, expm1, abs, mean,
#> sqrt, sign, round, concatenate, atleast_1d,
#> cos, sin, tan, cosh, sinh, tanh,
#> sum, prod, min, max, cumsum, cumprod )
#> ## note that above is from JAX numpy package, not numpy.
#>
#> y = np.array(r.carModelDF.getCard) #DATA
#> x = pd.factorize(np.array(r.carModelDF.carModel),use_na_sentinel=True)[0] #DIM
#> x_dim = len(np.unique(x)) #DIM
#> x_crd = pd.factorize(np.array(r.carModelDF.carModel),use_na_sentinel=True)[1] #DIM
#> def graph_model(y,x):
#> ## Define random variables and their relationships
#> with npo.plate('x_dim',x_dim):
#> theta = npo.sample('theta', dist.Beta(2,2))
#>
#> y = npo.sample('y', dist.Bernoulli(theta[x]),obs=y)
#>
#>
#> # computationally get posterior
#> mcmc = MCMC(NUTS(graph_model), num_warmup = 1000, num_samples = 4000)
#> rng_key = random.PRNGKey(seed = 1234567)
#> mcmc.run(rng_key,y,x)
#> axis_labels = {'x_dim': x_crd}
#> rv_to_axes = {'theta': ['x_dim']}
#> drop_names = set()
#> samples = mcmc.get_samples(group_by_chain=True) # dict: name -> [chains, draws, *axes]
#> import numpy as np, pandas as pd, string
#> # Flatten (chains*draws) and expand RVs using rv_to_axes + axis_labels
#> flat = {name: np.reshape(val, (-1,) + val.shape[2:]) for name, val in list(samples.items())}
#> out = {}
#> allowed = set(string.ascii_letters + string.digits + '._')
#> def sanitize_label(s):
#> s = str(s).strip().replace(' ', '.') # spaces -> dots
#> s = ''.join((ch if ch in allowed else '.') for ch in s)
#> # collapse repeated dots without regex
#> while '..' in s:
#> s = s.replace('..', '.')
#> s = s.strip('.')
#> return s if s else '1'
#>
#> for name, arr in list(flat.items()):
#> # Drop deterministic RVs entirely
#> if name in drop_names:
#> continue
#> axes = rv_to_axes.get(name, []) # [] if not present
#> # If scalar per draw, keep as a single column
#> if arr.ndim == 1:
#> out[name] = arr
#> continue
#> # Otherwise (vector/matrix/...): expand to separate columns
#> trailing = arr.shape[1:]
#> arr2 = arr.reshape(arr.shape[0], int(np.prod(trailing)))
#> for j in range(arr2.shape[1]):
#> idx = np.unravel_index(j, trailing)
#> parts = []
#> for axis, i in enumerate(idx):
#> if axis < len(axes):
#> axis_name = axes[axis]
#> labels = axis_labels.get(axis_name)
#> if labels is not None:
#> labs = np.asarray(labels).astype(str)
#> lab = labs[i] if i < labs.shape[0] else str(i + 1)
#> parts.append(sanitize_label(lab))
#> else:
#> parts.append(str(i + 1))
#> else:
#> parts.append(str(i + 1))
#> # No brackets; join labels with '_' so tibble won't append trailing dots
#> col = name if not parts else name + '_' + '_'.join(parts)
#> out[col] = arr2[:, j]
#> drawsDF = pd.DataFrame(out)"
#> ) ## END PYTHON STRING
#> drawsDF = reticulate::py$drawsDF
Getting Help and Suggesting Improvements
Whether you encounter a clear bug, have a suggestion for improvement, or just have a question, we are thrilled to help you out. In all cases, please file a GitHub issue. If reporting a bug, please include a minimal reproducible example.
Contributing
We welcome help turning causact into the most intuitive and fastest method of converting stakeholder narratives about data-generating processes into actionable insight from posterior distributions. If you want to help us achieve this vision, we welcome your contributions after reading the new contributor guide. Please note that this project is released with a Contributor Code of Conduct. By participating in this project you agree to abide by its terms.
Further Usage
For more info, see A Business Analyst's Introduction to Business Analytics available at https://www.causact.com. You can also check out the package’s vignette: vignette("narrative-to-insight-with-causact"). Two additional examples are shown below.
Prosocial Chimpanzees Example from Statistical Rethinking
McElreath, Richard. Statistical rethinking: A Bayesian course with examples in R and Stan. Chapman and Hall/CRC, 2018.
library(tidyverse)
library(causact)
# data object used below, chimpanzeesDF, is built-in to causact package
graph = dag_create() %>%
dag_node("Pull Left Handle","L",
rhs = bernoulli(p),
data = causact::chimpanzeesDF$pulled_left) %>%
dag_node("Probability of Pull", "p",
rhs = 1 / (1 + exp(-((alpha + gamma + beta)))),
child = "L") %>%
dag_node("Actor Intercept","alpha",
rhs = normal(alphaBar, sigma_alpha),
child = "p") %>%
dag_node("Block Intercept","gamma",
rhs = normal(0,sigma_gamma),
child = "p") %>%
dag_node("Treatment Intercept","beta",
rhs = normal(0,0.5),
child = "p") %>%
dag_node("Actor Population Intercept","alphaBar",
rhs = normal(0,1.5),
child = "alpha") %>%
dag_node("Actor Variation","sigma_alpha",
rhs = exponential(1),
child = "alpha") %>%
dag_node("Block Variation","sigma_gamma",
rhs = exponential(1),
child = "gamma") %>%
dag_plate("Observation","i",
nodeLabels = c("L","p")) %>%
dag_plate("Actor","act",
nodeLabels = c("alpha"),
data = chimpanzeesDF$actor,
addDataNode = TRUE) %>%
dag_plate("Block","blk",
nodeLabels = c("gamma"),
data = chimpanzeesDF$block,
addDataNode = TRUE) %>%
dag_plate("Treatment","trtmt",
nodeLabels = c("beta"),
data = chimpanzeesDF$treatment,
addDataNode = TRUE)
See graph
graph %>% dag_render(width = 2000, height = 800)
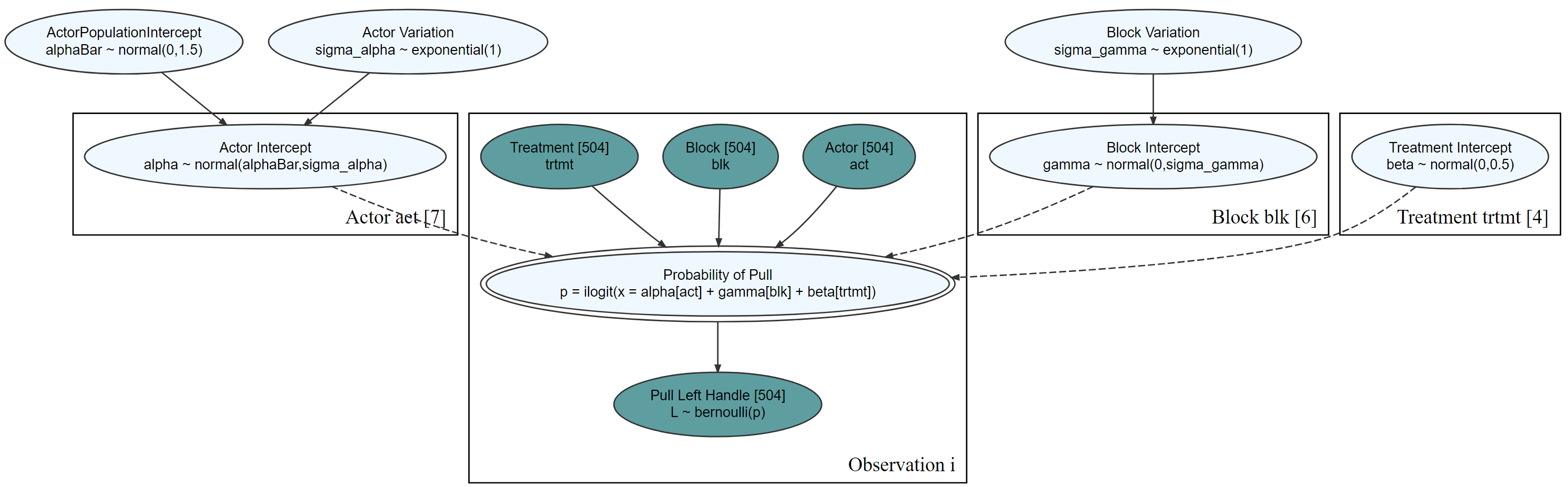
Communicate with stakeholders for whom the statistics might be distracting
graph %>% dag_render(shortLabel = TRUE)
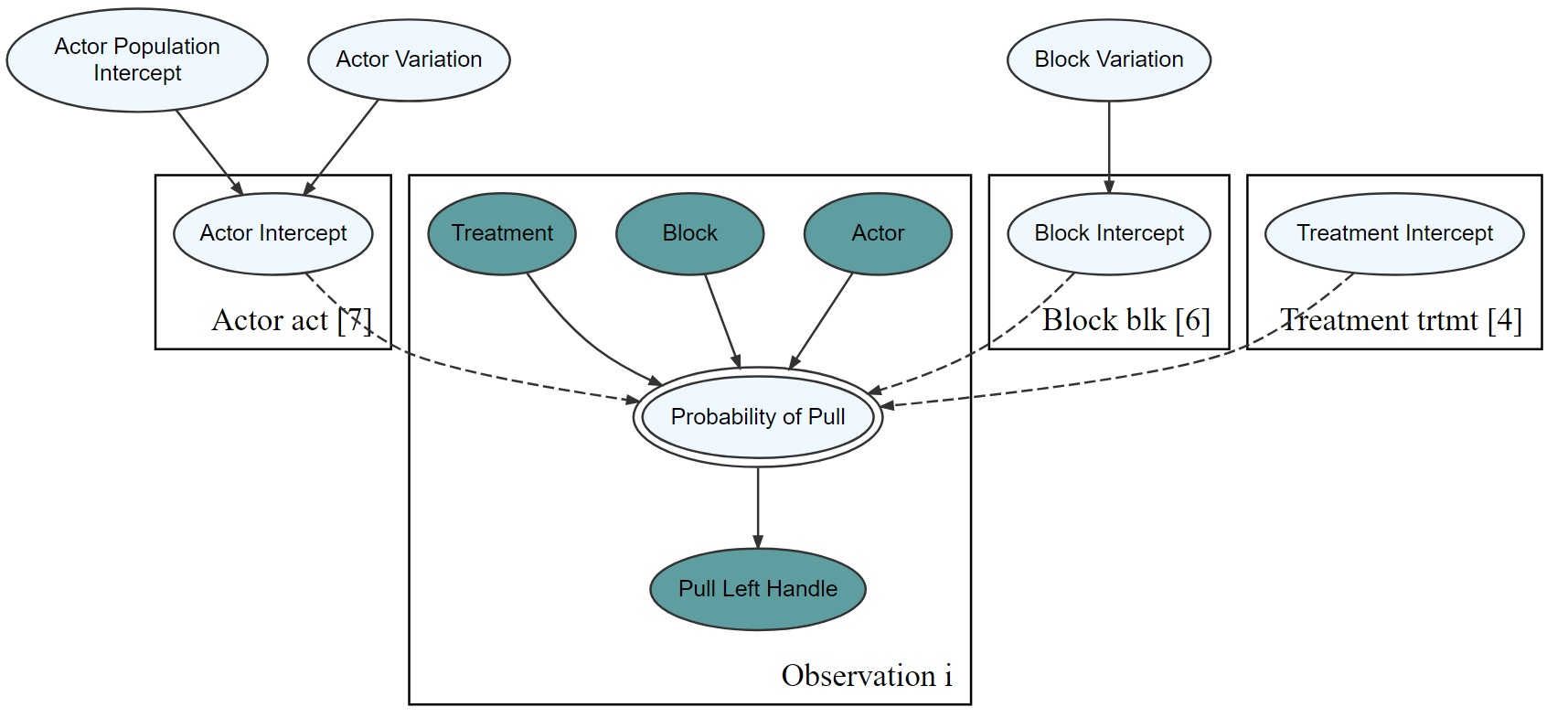
Compute posterior
drawsDF = graph %>% dag_numpyro()
Visualize posterior
drawsDF %>% dagp_plot()
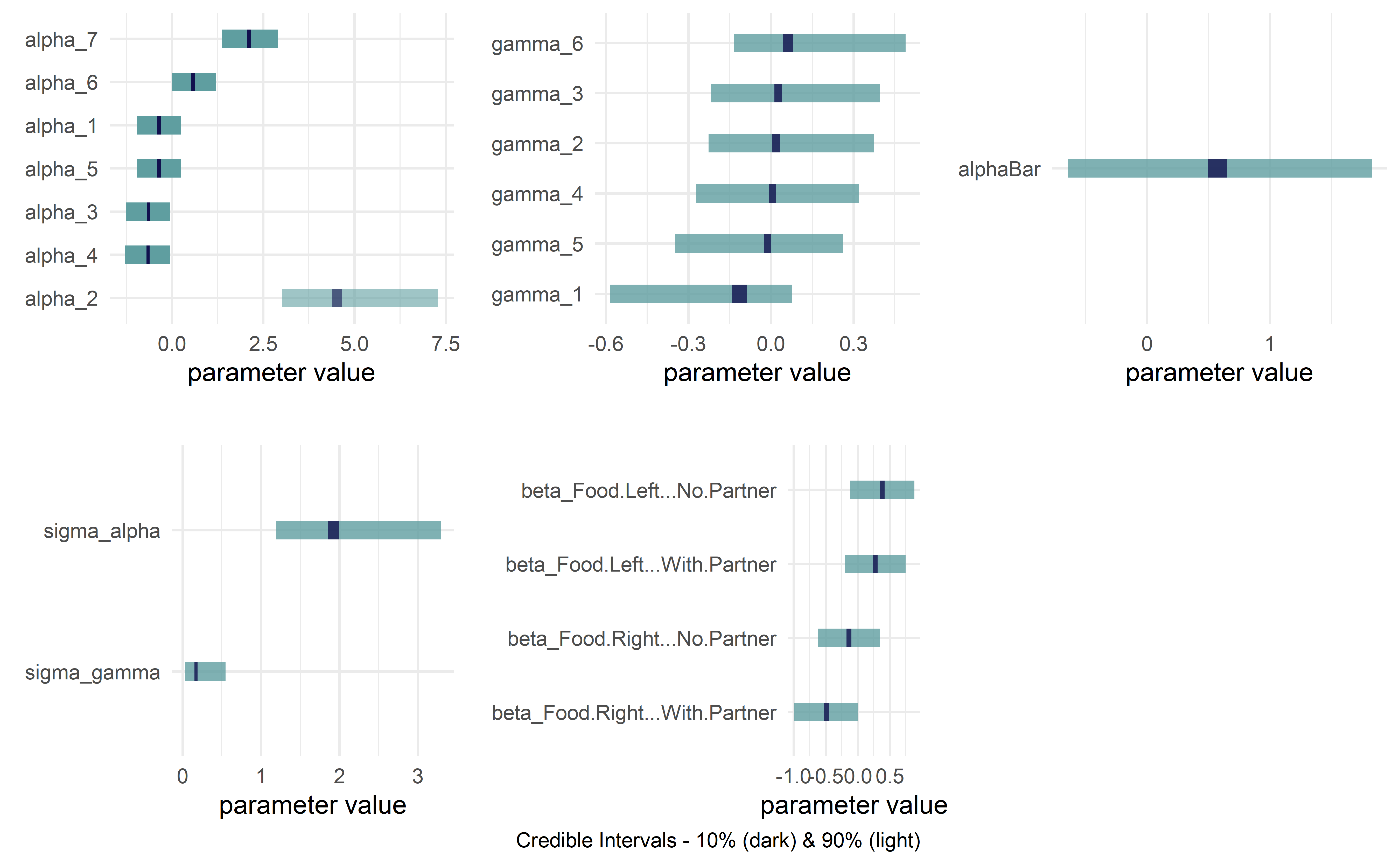
Eight Schools Example from Bayesian Data Analysis
Gelman, Andrew, Hal S. Stern, John B. Carlin, David B. Dunson, Aki Vehtari, and Donald B. Rubin. Bayesian data analysis. Chapman and Hall/CRC, 2013.
library(tidyverse)
library(causact)
# data object used below, schoolDF, is built-in to causact package
graph = dag_create() %>%
dag_node("Treatment Effect","y",
rhs = normal(theta, sigma),
data = causact::schoolsDF$y) %>%
dag_node("Std Error of Effect Estimates","sigma",
data = causact::schoolsDF$sigma,
child = "y") %>%
dag_node("Exp. Treatment Effect","theta",
child = "y",
rhs = avgEffect + schoolEffect) %>%
dag_node("Pop Treatment Effect","avgEffect",
child = "theta",
rhs = normal(0,30)) %>%
dag_node("School Level Effects","schoolEffect",
rhs = normal(0,30),
child = "theta") %>%
dag_plate("Observation","i",nodeLabels = c("sigma","y","theta")) %>%
dag_plate("School Name","school",
nodeLabels = "schoolEffect",
data = causact::schoolsDF$schoolName,
addDataNode = TRUE)
See graph
graph %>% dag_render()
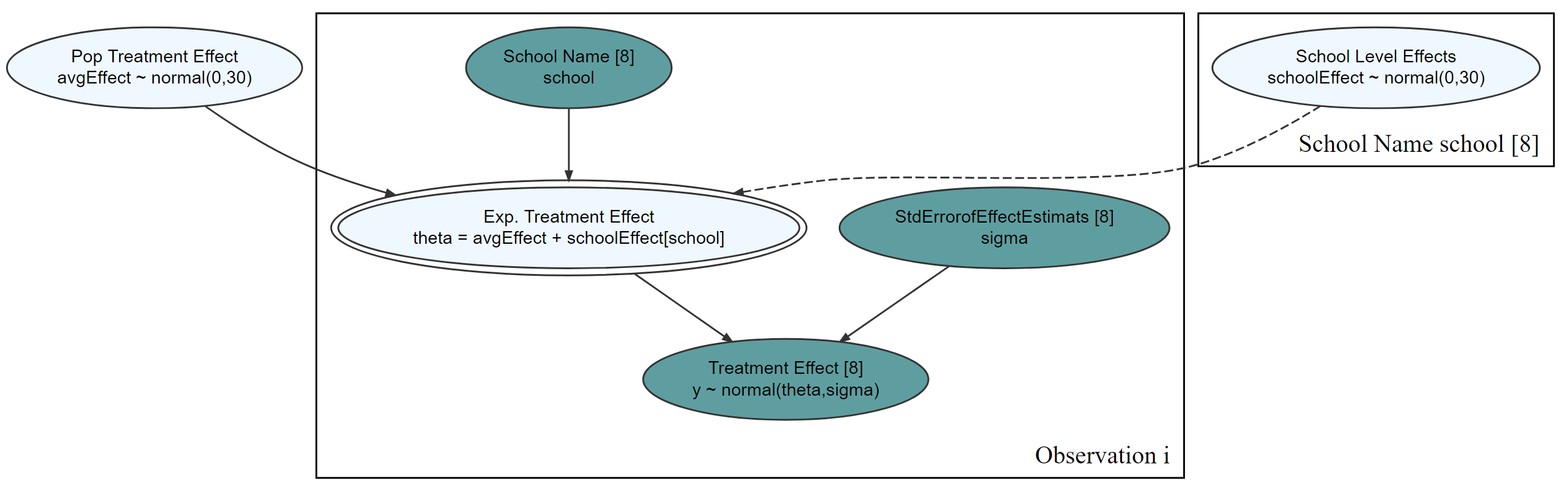
Compute posterior
drawsDF = graph %>% dag_numpyro()
Visualize posterior
drawsDF %>% dagp_plot()