Scientific Content and Citation Analysis from PDF Documents.
contentanalysis

Overview
contentanalysis is a comprehensive R package designed for in-depth analysis of scientific literature. It bridges the gap between raw PDF documents and structured, analyzable data by combining advanced text extraction, citation analysis, and bibliometric enrichment from external databases.
AI-Enhanced PDF Import: The package supports AI-assisted PDF text extraction through Google’s Gemini API, enabling more accurate parsing of complex document layouts. To use this feature, you need to obtain an API key from Google AI Studio.
Integration with bibliometrix: This package complements the science mapping analyses available in bibliometrix and its Shiny interface biblioshiny. If you want to perform content analysis within a user-friendly Shiny application with all the advantages of an interactive interface, simply install bibliometrix and launch biblioshiny, where you’ll find a dedicated Content Analysis menu that implements all the analyses and outputs of this library.
What Makes It Unique?
The package goes beyond simple PDF parsing by creating a multi-layered analytical framework:
Intelligent PDF Processing: Extracts text from multi-column PDFs while preserving document structure (sections, paragraphs, references)
Citation Intelligence: Detects and extracts citations in multiple formats (numbered, author-year, narrative, parenthetical) and maps them to their precise locations in the document
Bibliometric Enrichment: Automatically retrieves and integrates metadata from external sources:
- CrossRef API: Retrieves structured reference data including authors, publication years, journals, and DOIs
- OpenAlex: Enriches references with additional metadata, filling gaps and providing comprehensive bibliographic information
Citation-Reference Linking: Implements sophisticated matching algorithms to connect in-text citations with their corresponding references, handling various citation styles and ambiguous cases
Context-Aware Analysis: Extracts the textual context surrounding each citation, enabling semantic analysis of how references are used throughout the document
Network Visualization: Creates interactive networks showing citation co-occurrence patterns and conceptual relationships within the document
The Complete Workflow
PDF Document → Text Extraction → Citation Detection → Reference Parsing
↓
CrossRef/OpenAlex APIs
↓
Citation-Reference Matching → Enriched Dataset
↓
Network Analysis + Text Analytics + Bibliometric Indicators
The result is a rich, structured dataset that transforms a static PDF into an analyzable knowledge object, ready for: - Content analysis: Understanding what concepts and methods are discussed - Citation analysis: Examining how knowledge is constructed and referenced - Temporal analysis: Tracking the evolution of ideas through citation patterns - Network analysis: Visualizing intellectual connections - Readability assessment: Evaluating text complexity and accessibility
Key Features
PDF Import & Text Extraction
- Multi-column layout support with automatic section detection
- Structure preservation (title, abstract, introduction, methods, results, discussion, references)
- Handling of complex layouts and special characters
- DOI extraction from PDF metadata
Citation Extraction & Analysis
- Comprehensive detection of citation formats:
- Numbered citations:
[1],[1-3],[1,5,7]
- Numbered citations:
- Author-year citations:
(Smith, 2020),(Smith et al., 2020) - Narrative citations:
Smith (2020) demonstrated... - Complex citations:
(see Smith, 2020; Jones et al., 2021) - Citation context extraction (surrounding text analysis)
- Citation positioning and density metrics
- Section-wise citation distribution
Reference Management & Enrichment
- Local parsing: Extract references from the document’s reference section
- CrossRef integration: Retrieve structured metadata for cited works via DOI
- OpenAlex integration: Enrich references with additional bibliographic data
- Automatic gap-filling: Complete missing author names, years, journal names
- Structured reference format: Standardized author lists, publication years, journals
Citation-Reference Matching
- Intelligent matching algorithms with multiple confidence levels:
- High confidence: Exact author-year matches
- Medium confidence: Fuzzy matching for variant author names
- Disambiguation: Handles multiple works by the same author
- Support for various citation styles (APA, Chicago, Vancouver, etc.)
- Handles complex cases: multiple authors, “et al.”, year suffixes (2020a, 2020b)
Network Analysis
- Interactive citation co-occurrence networks
- Distance-based edge weighting (closer citations = stronger connections)
- Section-aware visualization (color-coded by document section)
- Multi-section citation detection (citations appearing in multiple sections)
- Network statistics: centrality, clustering, community detection potential
- Citation cluster descriptions: TF-IDF analysis of reference titles by section
- Interactive plotly visualizations: bar charts, heatmaps, and reference density plots
Text Analysis
- Word frequency analysis with stopword removal
- N-gram extraction (bigrams, trigrams)
- Lexical diversity metrics
- Readability indices (Flesch, Gunning Fog, SMOG, Coleman-Liau)
- Word distribution tracking across document sections
- Methodological term tracking
Bibliometric Indicators
- Citation density (citations per 1000 words)
- Citation type distribution (narrative vs. parenthetical)
- Co-citation analysis
- Reference age distribution
- Journal diversity metrics
Installation
You can install the development version from GitHub:
# install.packages("devtools")
devtools::install_github("massimoaria/contentanalysis")
Example
Complete workflow analyzing a real scientific paper:
library(contentanalysis)
Download example paper
The paper is an open access article by Aria et al.:
Aria, M., Cuccurullo, C., & Gnasso, A. (2021). A comparison among interpretative proposals for Random Forests. Machine Learning with Applications, 6, 100094.
paper_url <- "https://raw.githubusercontent.com/massimoaria/contentanalysis/master/inst/examples/example_paper.pdf"
download.file(paper_url, destfile = "example_paper.pdf", mode = "wb")
Import PDF with automatic section detection
doc <- pdf2txt_auto("example_paper.pdf",
n_columns = 2,
citation_type = "author_year")
#> Stripped running header (6 occurrences, 40 chars)
#> Using 17 sections from PDF table of contents
#> Found 16 sections: Preface, Introduction, Related work, Internal processing approaches, Random forest extra information, Visualization toolkits, Post-Hoc approaches, Size reduction, Rule extraction, Local explanation, Comparison study, Experimental design, Analysis, Conclusion, Acknowledgment, References
#> Normalized 77 references with consistent \n\n separators
# Check detected sections
names(doc)
#> [1] "Full_text" "Preface"
#> [3] "Introduction" "Related work"
#> [5] "Internal processing approaches" "Random forest extra information"
#> [7] "Visualization toolkits" "Post-Hoc approaches"
#> [9] "Size reduction" "Rule extraction"
#> [11] "Local explanation" "Comparison study"
#> [13] "Experimental design" "Analysis"
#> [15] "Conclusion" "Acknowledgment"
#> [17] "References"
Perform comprehensive content analysis with CrossRef and OpenAlex enrichment
analysis <- analyze_scientific_content(
text = doc,
doi = "10.1016/j.mlwa.2021.100094",
mailto = "[email protected]",
citation_type = "author_year"
)
#> Extracting author-year citations only
#> Attempting to retrieve references from CrossRef...
#> Successfully retrieved 33 references from CrossRef
#> Fetching Open Access metadata for 14 DOIs from OpenAlex...
#> Successfully retrieved metadata for 13 references from OpenAlex
This single function call:
- Extracts all citations from the document
- Retrieves reference metadata from CrossRef using the paper’s DOI
- Enriches references with additional data from OpenAlex
- Matches citations to references with confidence scoring
- Performs text analysis and computes bibliometric indicators
View summary statistics
analysis$summary
#> $total_words_analyzed
#> [1] 3453
#>
#> $unique_words
#> [1] 1310
#>
#> $citations_extracted
#> [1] 49
#>
#> $narrative_citations
#> [1] 15
#>
#> $parenthetical_citations
#> [1] 34
#>
#> $complex_citations_parsed
#> [1] 12
#>
#> $lexical_diversity
#> [1] 0.3793802
#>
#> $average_citation_context_length
#> [1] 3230.429
#>
#> $citation_density_per_1000_words
#> [1] 6.47
#>
#> $references_parsed
#> [1] 33
#>
#> $citations_matched_to_refs
#> [1] 42
#>
#> $match_quality
#> # A tibble: 3 × 3
#> match_confidence n percentage
#> <chr> <int> <dbl>
#> 1 high 42 85.7
#> 2 no_match_author 6 12.2
#> 3 no_match_year 1 2
#>
#> $citation_type_used
#> [1] "author_year"
Readability indices
readability <- calculate_readability_indices(doc$Full_text, detailed = TRUE)
readability
#> # A tibble: 1 × 12
#> flesch_kincaid_grade flesch_reading_ease automated_readability_index
#> <dbl> <dbl> <dbl>
#> 1 12.7 33.3 12.1
#> # ℹ 9 more variables: gunning_fog_index <dbl>, n_sentences <int>,
#> # n_words <int>, n_syllables <dbl>, n_characters <int>,
#> # n_complex_words <int>, avg_sentence_length <dbl>,
#> # avg_syllables_per_word <dbl>, pct_complex_words <dbl>
Examine citations by type
analysis$citation_metrics$type_distribution
#> # A tibble: 11 × 3
#> citation_type n percentage
#> <chr> <int> <dbl>
#> 1 parsed_from_multiple 12 24.5
#> 2 author_year_basic 9 18.4
#> 3 narrative_etal 7 14.3
#> 4 author_year_and 6 12.2
#> 5 author_year_etal 3 6.12
#> 6 narrative_three_authors_and 3 6.12
#> 7 narrative_two_authors_and 3 6.12
#> 8 narrative_four_authors_and 2 4.08
#> 9 see_citations 2 4.08
#> 10 author_year_ampersand 1 2.04
#> 11 doi_pattern 1 2.04
Analyze citation contexts
head(analysis$citation_contexts[, c("citation_text_clean", "section", "full_context")])
#> # A tibble: 6 × 3
#> citation_text_clean section full_context
#> <chr> <chr> <chr>
#> 1 (Mitchell, 1997) Introd… systems ide…
#> 2 https://doi.org/10.1016/j.mlwa.2021.100094 Introd… interpretat…
#> 3 (Breiman, 2001) Introd… random subs…
#> 4 (see Breiman, 1996) Introd… model that …
#> 5 (Hastie, supervised learning (Breiman, Friedman, Tibshir… Introd… by calculat…
#> 6 (Hastie et al., 2009) Introd… but it is n…
Citation Network Visualization
Create interactive network visualizations showing how citations co-occur within your document:
# Create citation network
network <- create_citation_network(
citation_analysis_results = analysis,
max_distance = 800, # Max distance between citations (characters)
min_connections = 2, # Minimum connections to include a node
show_labels = TRUE
)
# Display interactive network
network
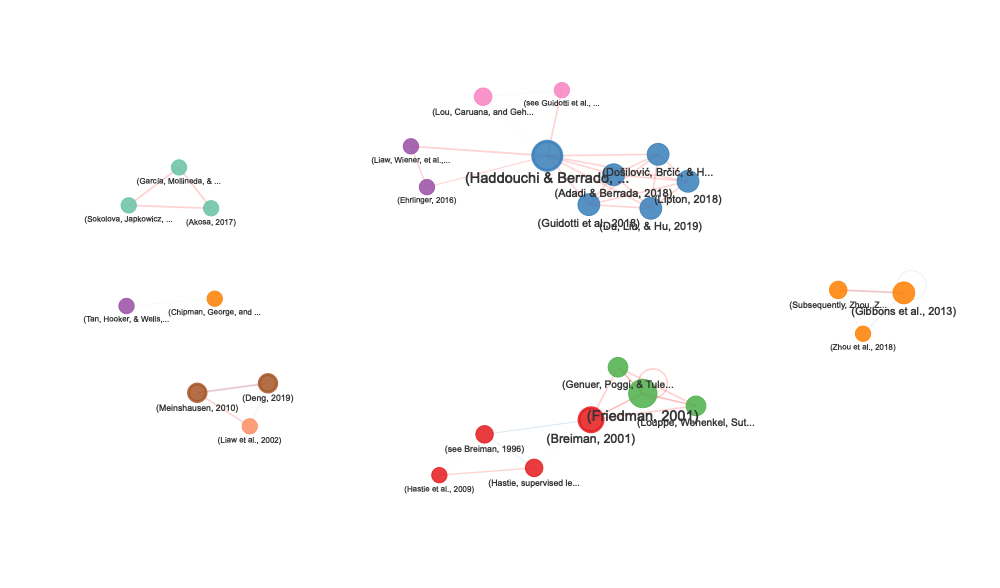
Access network statistics
stats <- attr(network, "stats")
# Network size
cat("Nodes:", stats$n_nodes, "\n")
#> Nodes: 28
cat("Edges:", stats$n_edges, "\n")
#> Edges: 49
cat("Average distance:", stats$avg_distance, "characters\n")
#> Average distance: 245.4 characters
# Citations by section
print(stats$section_distribution)
#> primary_section n
#> 1 Related work 6
#> 2 Introduction 4
#> 3 Size reduction 4
#> 4 Experimental design 3
#> 5 Random forest extra information 3
#> 6 Visualization toolkits 3
#> 7 Local explanation 2
#> 8 Rule extraction 2
#> 9 Analysis 1
# Multi-section citations
if (nrow(stats$multi_section_citations) > 0) {
print(stats$multi_section_citations)
}
#> citation_text
#> 1 (Breiman, 2001)
#> 2 (Haddouchi & Berrado, 2019)
#> 3 (Meinshausen, 2010)
#> 4 (Deng, 2019)
#> sections n_sections
#> 1 Introduction, Random forest extra information 2
#> 2 Related work, Random forest extra information, Rule extraction 3
#> 3 Rule extraction, Comparison study, Analysis 3
#> 4 Rule extraction, Comparison study, Analysis 3
Network Features
The citation network visualization includes:
- Node size: Proportional to number of connections
- Node color: Indicates the primary section where citations appear
- Node border: Thicker border (3px) for citations appearing in multiple sections
- Edge thickness: Decreases with distance (closer citations = thicker edges)
- Edge color:
- Red: Very close citations (≤300 characters)
- Blue: Moderate distance (≤600 characters)
- Gray: Distant citations (>600 characters)
- Interactive features: Zoom, pan, drag nodes, highlight neighbors on hover
Customizing the Network
# Focus on very close citations only
network_close <- create_citation_network(
analysis,
max_distance = 300,
min_connections = 1
)
# Show only highly connected citations
network_hubs <- create_citation_network(
analysis,
max_distance = 1000,
min_connections = 5
)
# Hide labels for cleaner visualization
network_clean <- create_citation_network(
analysis,
show_labels = FALSE
)
Citation Cluster Description
Describe the thematic focus of each section’s bibliography using TF-IDF analysis of reference titles:
# Generate cluster descriptions
cluster_desc <- describe_citation_clusters(analysis, top_n = 10)
# View summary: top terms per section
cluster_desc$cluster_summary
#> # A tibble: 10 × 3
#> section n_references top_terms
#> <chr> <int> <chr>
#> 1 Introduction 4 learning, machine, machine lear…
#> 2 Related work 8 black, black box, box, acm, sur…
#> 3 Random forest extra information 5 forests, random forests, annals…
#> 4 Visualization toolkits 4 analytics, analytics ieee, comp…
#> 5 Size reduction 4 adaptive, adaptive diagnostic, …
#> 6 Rule extraction 2 annals applied, applied, applie…
#> 7 Local explanation 3 models, classification, classif…
#> 8 Comparison study 1 annals applied, applied, applie…
#> 9 Experimental design 3 bell, bell laboratories, labora…
#> 10 Analysis 3 domains, domains acm, imbalance…
# View detailed TF-IDF scores
cluster_desc$cluster_descriptions
#> # A tibble: 83 × 7
#> section ngram ngram_size n tf idf tf_idf
#> <chr> <chr> <int> <int> <dbl> <dbl> <dbl>
#> 1 Introduction learning 1 2 0.143 1.20 0.172
#> 2 Introduction machine 1 2 0.143 1.20 0.172
#> 3 Introduction machine learning 2 2 0.143 1.20 0.172
#> 4 Introduction bagging 1 1 0.0714 2.30 0.164
#> 5 Introduction bagging predictors 2 1 0.0714 2.30 0.164
#> 6 Introduction predictors 1 1 0.0714 2.30 0.164
#> 7 Introduction predictors machine 2 1 0.0714 2.30 0.164
#> 8 Introduction forests machine 2 1 0.0714 1.61 0.115
#> 9 Introduction forests 1 1 0.0714 1.20 0.0860
#> 10 Introduction random forests 2 1 0.0714 1.20 0.0860
#> # ℹ 73 more rows
Visualize Citation Clusters
Create interactive plotly visualizations that complement the citation network:
1. TF-IDF terms per section (2-column grid layout)
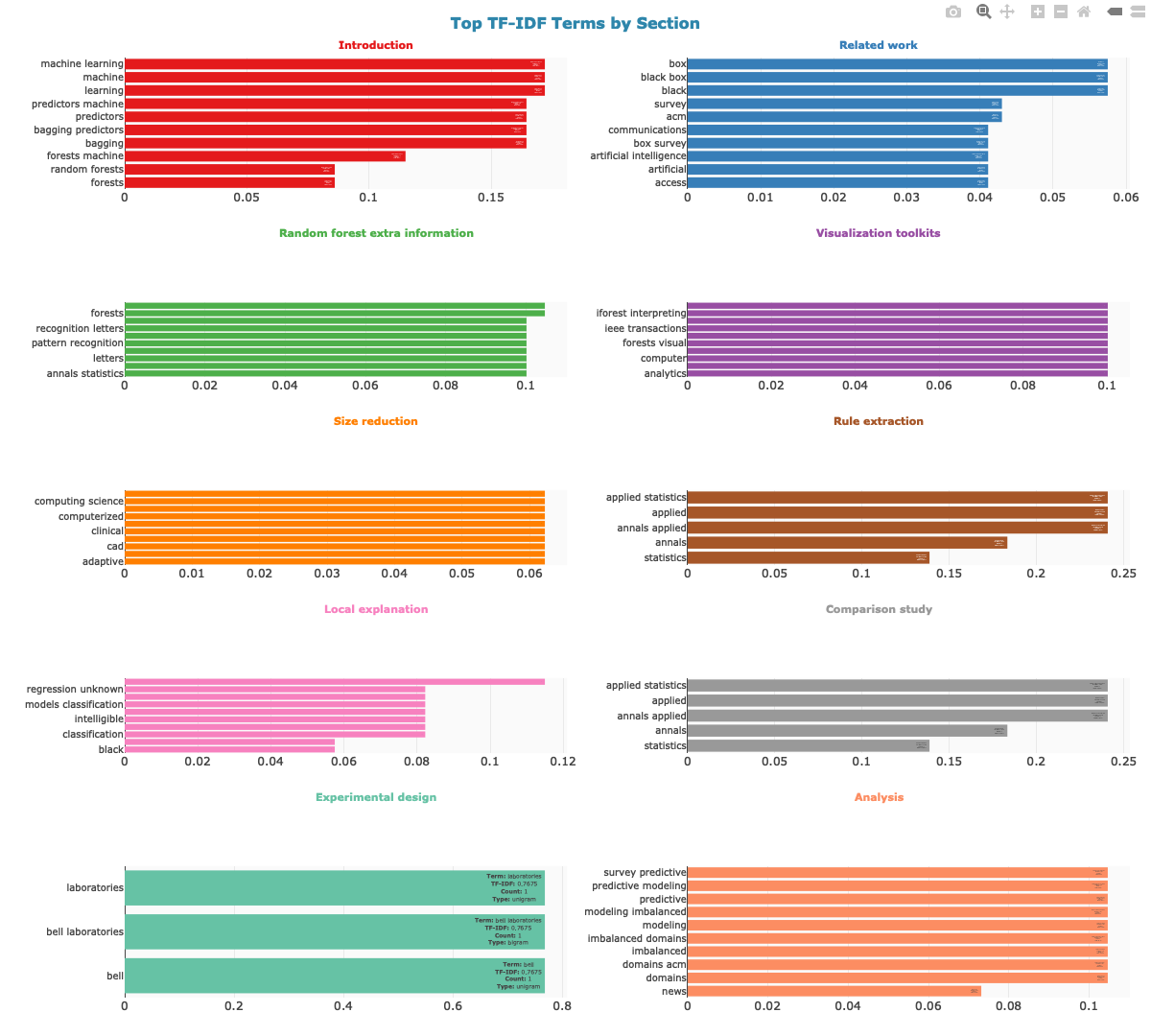
2. Heatmap: terms vs sections (unique vs shared terms)
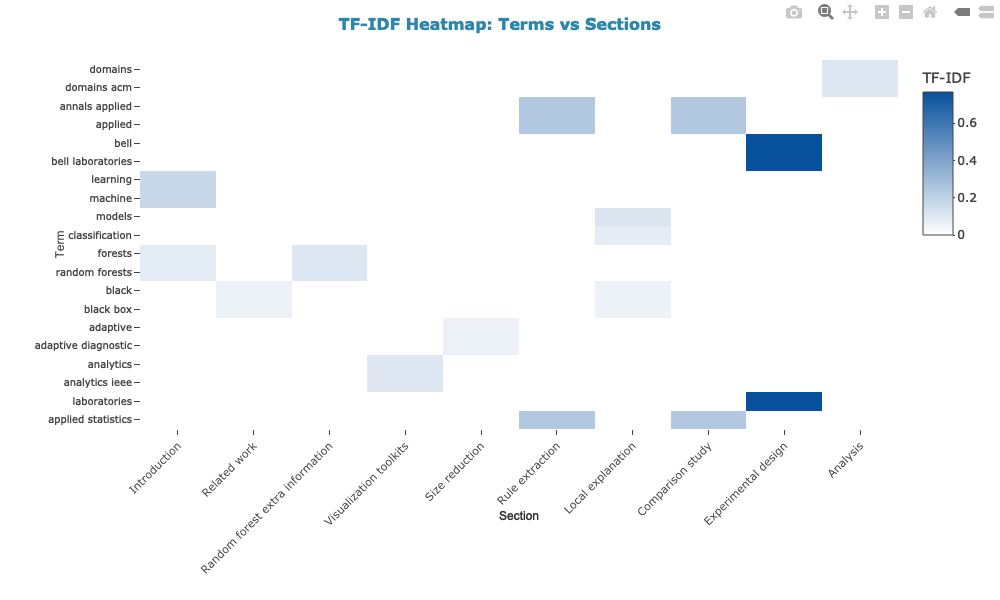
3. References per section
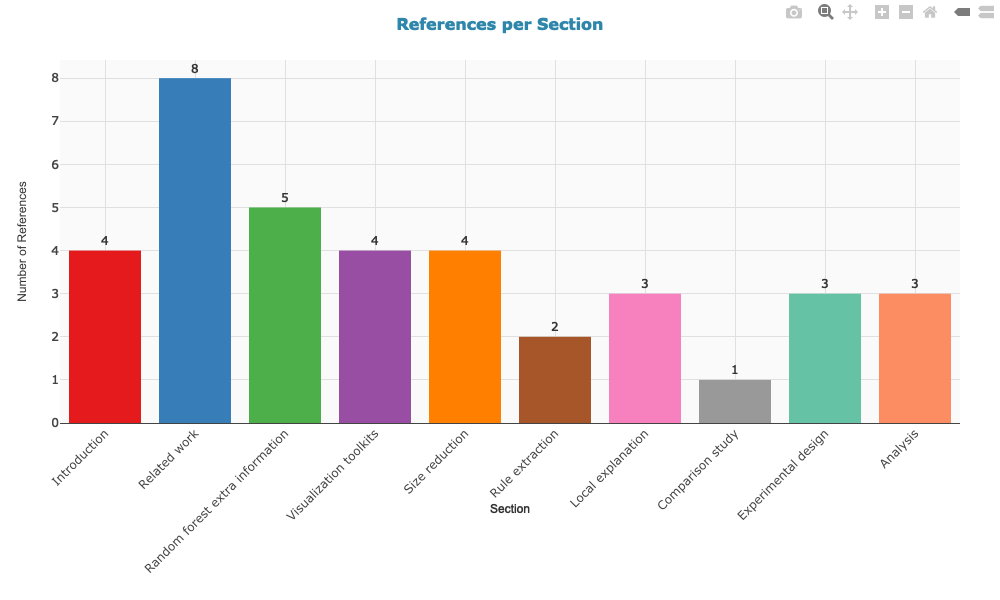
The TF-IDF bar chart uses a 2-column grid layout with color-coded section titles for a compact, readable overview. All plots display sections in the order they appear in the paper, use consistent styling, and include interactive hover information. Colors match the section colors used in the citation network.
Text Analysis
Track methodological terms across sections
method_terms <- c("machine learning", "regression", "validation", "dataset")
word_dist <- calculate_word_distribution(doc, method_terms)
Create interactive visualization
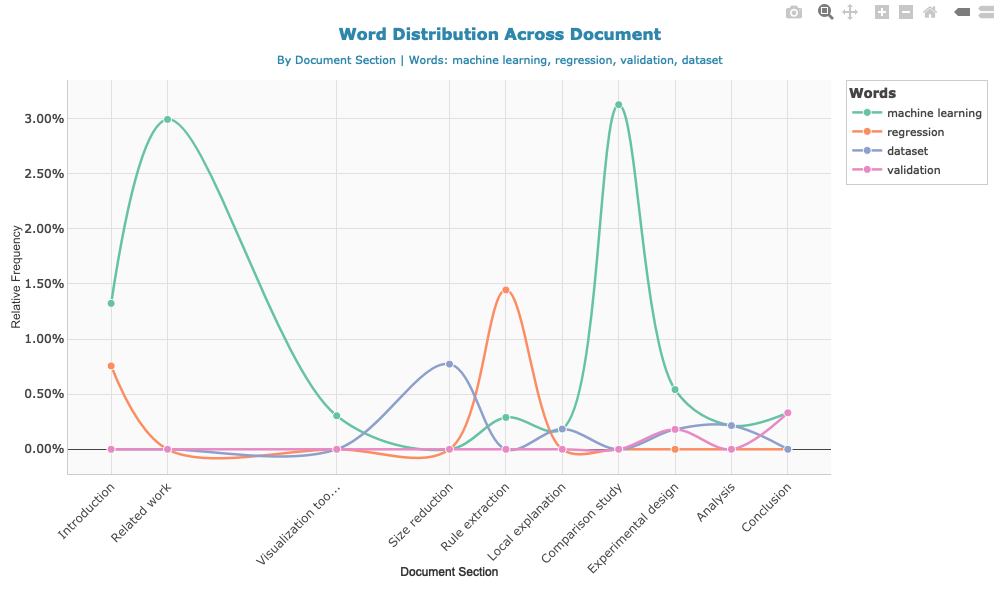
Examine most frequent words
head(analysis$word_frequencies, 10)
#> # A tibble: 10 × 4
#> word n frequency rank
#> <chr> <int> <dbl> <int>
#> 1 model 45 0.0130 1
#> 2 forest 42 0.0122 2
#> 3 accuracy 40 0.0116 3
#> 4 trees 38 0.0110 4
#> 5 random 34 0.00985 5
#> 6 learning 27 0.00782 6
#> 7 set 27 0.00782 7
#> 8 variable 26 0.00753 8
#> 9 data 25 0.00724 9
#> 10 rule 25 0.00724 10
Citation co-occurrence data
head(analysis$network_data)
#> # A tibble: 6 × 5
#> citation1 citation2 distance type1 type2
#> <chr> <chr> <int> <chr> <chr>
#> 1 (Mitchell, 1997) https://… 734 auth… doi_…
#> 2 (Breiman, 2001) (see Bre… 318 auth… see_…
#> 3 (Breiman, 2001) (Hastie,… 734 auth… auth…
#> 4 (see Breiman, 1996) (Hastie,… 398 see_… auth…
#> 5 (see Breiman, 1996) (Hastie … 685 see_… auth…
#> 6 (Hastie, supervised learning (Breiman, Friedma… (Hastie … 210 auth… auth…
Working with References
Exploring enriched reference data
# View parsed references (enriched with CrossRef and OpenAlex)
head(analysis$parsed_references[, c("ref_first_author", "ref_year", "ref_journal", "ref_source")])
#> ref_first_author ref_year ref_journal ref_source
#> 1 Adadi 2018 IEEE Access crossref
#> 2 <NA> <NA> <NA> crossref
#> 3 Branco 2016 ACM Computing Surveys crossref
#> 4 Breiman 1996 Machine Learning crossref
#> 5 Breiman 2001 Machine Learning crossref
#> 6 Breiman 1984 International Group crossref
# Check data sources
table(analysis$parsed_references$ref_source)
#>
#> crossref
#> 33
The ref_source column indicates where the data came from:
"crossref": Retrieved from CrossRef API"parsed": Extracted from document’s reference section- References may be enriched with OpenAlex data even if originally from CrossRef
Analyzing citation-reference links
# View citation-reference matches with confidence levels
library(dplyr)
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
head(analysis$citation_references_mapping[, c("citation_text_clean", "ref_authors",
"ref_year", "match_confidence")])
#> # A tibble: 6 × 4
#> citation_text_clean ref_authors ref_year match_confidence
#> <chr> <chr> <chr> <chr>
#> 1 (Mitchell, 1997) Mitchell 1997 high
#> 2 https://doi.org/10.1016/j.mlwa.2021.100… <NA> <NA> no_match_year
#> 3 (Breiman, 2001) Breiman, L. 2001 high
#> 4 (see Breiman, 1996) Breiman, L. 1996 high
#> 5 (Hastie, supervised learning (Breiman, … Hastie 2009 high
#> 6 (Hastie et al., 2009) Hastie 2009 high
# Match quality distribution
table(analysis$citation_references_mapping$match_confidence)
#>
#> high no_match_author no_match_year
#> 42 6 1
Finding citations to specific authors
# Find all citations to works by Smith
analysis$citation_references_mapping %>%
filter(grepl("Smith", ref_authors, ignore.case = TRUE)) %>%
select(citation_text_clean, ref_full_text, match_confidence)
#> # A tibble: 0 × 3
#> # ℹ 3 variables: citation_text_clean <chr>, ref_full_text <chr>,
#> # match_confidence <chr>
Accessing OpenAlex metadata
# If OpenAlex data was retrieved
if (!is.null(analysis$references_oa)) {
# View enriched metadata
head(analysis$references_oa[, c("title", "publication_year", "cited_by_count",
"type", "oa_status")])
# Analyze citation impact
summary(analysis$references_oa$cited_by_count)
}
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> 100 205 1056 12429 5399 119748
Citations by section
analysis$citation_metrics$section_distribution
#> # A tibble: 15 × 3
#> section n percentage
#> <fct> <int> <dbl>
#> 1 Preface 0 0
#> 2 Introduction 6 12.2
#> 3 Related work 9 18.4
#> 4 Internal processing approaches 0 0
#> 5 Random forest extra information 6 12.2
#> 6 Visualization toolkits 4 8.16
#> 7 Post-Hoc approaches 0 0
#> 8 Size reduction 6 12.2
#> 9 Rule extraction 3 6.12
#> 10 Local explanation 5 10.2
#> 11 Comparison study 2 4.08
#> 12 Experimental design 4 8.16
#> 13 Analysis 4 8.16
#> 14 Conclusion 0 0
#> 15 Acknowledgment 0 0
Advanced: Word Distribution Analysis
# Track disease-related terms
disease_terms <- c("covid", "pandemic", "health", "policy", "vaccination")
dist <- calculate_word_distribution(doc, disease_terms, use_sections = TRUE)
# View frequencies by section
dist %>%
select(segment_name, word, count, percentage) %>%
arrange(segment_name, desc(percentage))
#> # A tibble: 1 × 4
#> segment_name word count percentage
#> <chr> <chr> <int> <dbl>
#> 1 Conclusion health 1 0.331
# Visualize trends
#plot_word_distribution(dist, plot_type = "area", smooth = FALSE)
Main Functions
PDF Import
pdf2txt_auto(): Import PDF with automatic section detectionreconstruct_text_structured(): Advanced text reconstructionextract_doi_from_pdf(): Extract DOI from PDF metadata
Content Analysis
analyze_scientific_content(): Comprehensive content and citation analysis with API enrichmentparse_references_section(): Parse reference list from textmatch_citations_to_references(): Match citations to references with confidence scoringget_crossref_references(): Retrieve references from CrossRef API
Network Analysis
create_citation_network(): Create interactive citation co-occurrence networkdescribe_citation_clusters(): Describe citation clusters by section using TF-IDF on reference titlesplot_citation_clusters(): Create interactive plotly visualizations of citation cluster descriptions (TF-IDF bars, heatmap, references per section)
Text Analysis
calculate_readability_indices(): Compute readability scores (Flesch, Gunning Fog, SMOG, Coleman-Liau)calculate_word_distribution(): Track word frequencies across document sectionsreadability_multiple(): Batch readability analysis for multiple documents
Visualization
plot_word_distribution(): Interactive visualization of word distribution across sectionsplot_citation_clusters(): Interactive TF-IDF bar charts, heatmaps, and reference density plots
Utilities
get_example_paper(): Download example paper for testingmap_citations_to_segments(): Map citations to document sections/segments
External Data Sources
CrossRef API
The package integrates with CrossRef’s REST API to retrieve structured bibliographic data:
- Endpoint:
https://api.crossref.org/works/{doi}/references - Data retrieved: Authors, publication year, journal/source, article title, DOI
- Rate limits: Polite pool requires email (use
mailtoparameter) - More info: https://api.crossref.org
OpenAlex API
OpenAlex provides comprehensive scholarly metadata:
- Endpoint: Via
openalexRpackage - Data retrieved: Complete author lists, citation counts, open access status, institutional affiliations
- Rate limits: 100,000 requests/day (polite pool with email), 10 requests/second
- API key: Optional, increases rate limits. Set with
openalexR::oa_apikey() - More info: https://openalex.org
Setting Up API Access
Email for polite pool (recommended)
Both CrossRef and OpenAlex offer a polite pool for users who provide an email address. This gives you faster and more reliable access compared to anonymous requests. Simply pass your email via the mailto parameter:
analysis <- analyze_scientific_content(
text = doc,
doi = "10.xxxx/xxxxx",
mailto = "[email protected]" # Your email for polite pool access
)
The mailto parameter is used by both CrossRef and OpenAlex to route your requests through their polite pool, which provides higher rate limits and priority access.
OpenAlex API key (optional, for extended use)
For heavier usage, OpenAlex offers a free API key that further increases rate limits (from 10 to 100 requests/second). This is recommended if you plan to analyze many documents in batch.
- Get your free API key at: https://openalex.org/users
- Set it in your R session before running the analysis:
openalexR::oa_apikey("your-api-key-here")
You can also add this to your .Rprofile so it’s automatically set at startup:
# Add to ~/.Rprofile
openalexR::oa_apikey("your-api-key-here")
Dependencies
Core: pdftools, dplyr, tidyr, stringr, tidytext, tibble, httr2, visNetwork, openalexR
Suggested: plotly, RColorBrewer, scales (for visualization)
Citation
If you use this package in your research, please cite:
Massimo Aria & Corrado Cuccurullo (2025). contentanalysis: Scientific Content and Citation Analysis from PDF Documents.
R package version 0.2.0.
https://doi.org/10.32614/CRAN.package.contentanalysis
License
GPL (>= 3)
Issues and Contributions
Please report issues at: https://github.com/massimoaria/contentanalysis/issues
Contributions are welcome! Please feel free to submit a Pull Request.