Description
CURE (Cumulative Residual) Plots.
Description
Creates 'ggplot2' Cumulative Residual (CURE) plots to check the goodness-of-fit of a count model; or the tables to create a customized version. A dataset of crashes in Washington state is available for illustrative purposes.
README.md
cureplots
Cumulative residual (CURE) plots assess the goodness-of-fit of a covariate in a generalized linear regression model, usually a negative binomial regression or a Poisson regression. The package cureplots produces CURE plots for the requested variables produced with ggplot2 or a table to easily produce a customized plot with the desired package.
Installation
Install the latest CRAN version with
install.packages("cureplots")
You can install the development version of cureplots from GitHub with the following:
# install.packages("devtools")
devtools::install_github("gbasulto/cureplots")
Functions and Data
| Name | Purpose |
|---|---|
calculate_cure_dataframe | Calculate CURE dataframe. Useful to produce customized CURE plots or CURE plots with variables not included in the model or transformations of variables that were included in the model (e.g., CURE plot for AADT when log(AADT) was included in the model). |
cure_plot | Produce default CURE plot by either providing model and variable to plot or an output from calculate_cure_dataframe function. |
resample_residuals | Resample cumulative residuals to overlay to CURE plots and better interpret results. |
washington_roads | Curated dataframe of crashes in Washington roads. |
Functions in cureplots
Basic Example
The example below shows
- How to produce a cure plot directly from the model object and
- How to produce the table to customize a plot.
A Poisson GLM model is adjusted to simulated data using the package glm. The functions also work with the gam package.
library(cureplots)
## basic example
set.seed(2000)
## Define parameters
beta <- c(-1, 0.3, 3)
## Simulate idependent variables
n <- 900
AADT <- c(runif(n, min = 2000, max = 150000))
nlanes <- sample(x = c(2, 3, 4), size = n, replace = TRUE)
LNAADT <- log(AADT)
## Simulate dependent variable
theta <- exp(beta[1] + beta[2] * LNAADT + beta[3] * nlanes)
y <- rpois(n, theta)
## Fit model
mod <- glm(y ~ LNAADT + nlanes, family = poisson)
## Calculate residuals
res <- residuals(mod, type = "response")
## Calculate CURE plot data
cure_df <- calculate_cure_dataframe(AADT, res)
#> Covariate: AADT
head(cure_df)
#> # A tibble: 6 × 5
#> AADT residual cumres lower upper
#> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 2363. -233. -233. -457. 457.
#> 2 2435. 17.2 -216. -459. 459.
#> 3 2724. 246. 29.9 -666. 666.
#> 4 2978. -1539. -1509. -3081. 3081.
#> 5 3007. -19.5 -1528. -3081. 3081.
#> 6 3149. -338. -1867. -3151. 3151.
## Providing CURE data frame
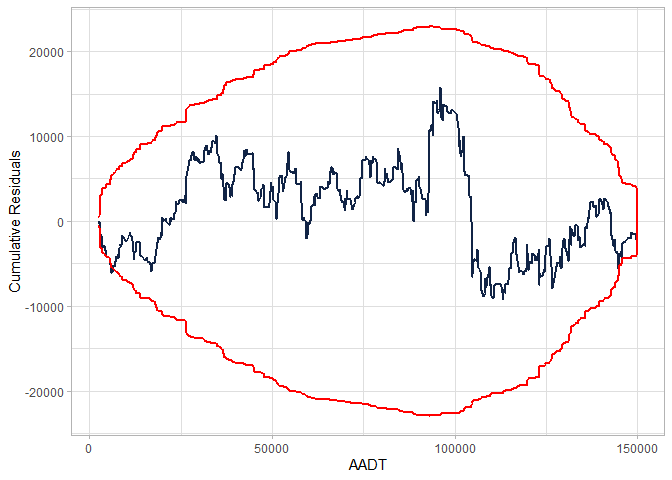
cure_plot(cure_df)
#> CURE data frame was provided. Its first column, AADT, will be used.
## Providing glm object
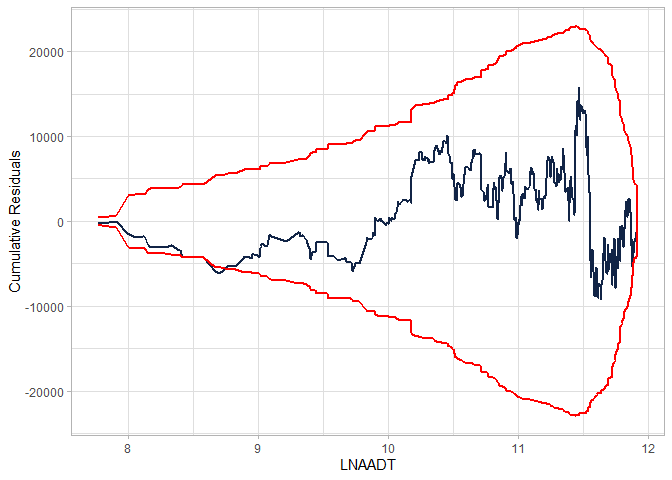
cure_plot(mod, "LNAADT")
#> Covariate LNAADT will be used to produce CURE plot.
Example with Resampling
library(cureplots)
## Basic example
set.seed(2000)
## Define parameters.
beta <- c(-1, 0.3, 3)
## Simulate idependent variables
n <- 900
AADT <- c(runif(n, min = 2000, max = 150000))
nlanes <- sample(x = c(2, 3, 4), size = n, replace = TRUE)
LNAADT <- log(AADT)
## Simulate dependent variable
theta <- exp(beta[1] + beta[2] * LNAADT + beta[3] * nlanes)
y <- rpois(n, theta)
## Fit model
mod <- glm(y ~ LNAADT + nlanes, family = poisson)
## Calculate residuals
res <- residuals(mod, type = "response")
## Calculate CURE plot data
cure_df <- calculate_cure_dataframe(AADT, res)
#> Covariate: AADT
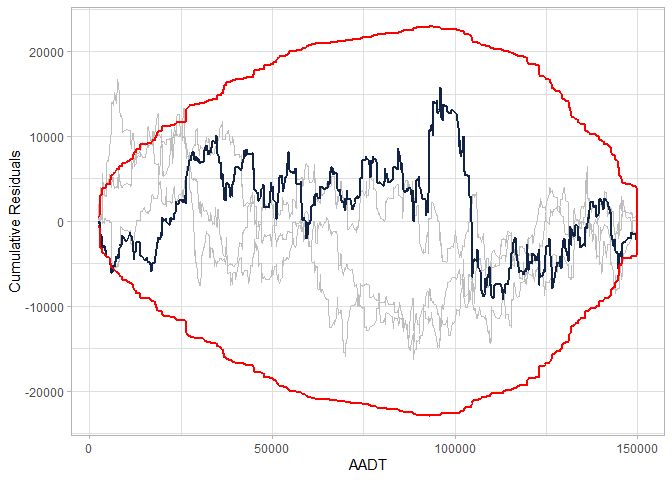
cure_plot(cure_df, n_resamples = 3)
#> CURE data frame was provided. Its first column, AADT, will be used.