Utilities for Joining Dataframes with Inexact Matching.
fozziejoin 🧸
⚠️ Note: This is a new R package, not yet on CRAN. Installation requires the Rust toolchain.
fozziejoin is an R package that performs fast fuzzy joins using Rust as a backend. It is a performance-minded re-imagining of the very popular fuzzyjoin package. Performance improvements relative to fuzzyjoin can be significant, especially for string distance joins. See the benchmarks for more details.
Currently, the following function families are available:
fozzie_string_joinfozzie_difference_joinfozzie_distance_joinfozzie_interval_joinfozzie_interval_joinfozzie_regex_joinfozzie_temporal_joinfozzie_temporal_interval_join
These function families include related functions, such as fozzie_string_inner_join.
The name is a playful nod to “fuzzy join” — reminiscent of Fozzie Bear from the Muppets. A picture of Fozzie will appear in the repo once the legal team gets braver. Wocka wocka!
Requirements
R 4.2 or greater is required for all installations. R 4.5.0 or greater is preferred.
On Linux or to build from source, you will need these additional dependencies:
- Cargo, the Rust package manager
- Rustc
- xz
While note strictly required, many of the installation instructions assume devtools is installed.
To run the examples in the README or benchmarking scripts, the following are required:
dplyrfuzzyjoinqdapDictionariesmicrobenchmarktibble
Installation
fozziejoin is currently under development for a future CRAN release. Until CRAN acceptance, installing from source is the only option. An appropriate Rust toolchain is required.
Linux/MacOS
devtools::install_github("fozzieverse/fozziejoin/fozziejoin-r")
Windows
To compile Rust extensions for R on Windows (such as those used by rextendr), you must use the GNU Rust toolchain, not MSVC. This is because R is built with GCC (via Rtools), and Rust must match that ABI for compatibility. This assumes you already have Rust installed.
- Set the default Rust toolchain to GNU:
# Install the GNU toolchain if needed
# rustup install stable-x86_64-pc-windows-gnu
rustup override set stable-x86_64-pc-windows-gnu
- Install the latest build from GitHub
Rscript -e 'devtools::install_github("fozzieverse/fozziejoin/fozziejoin-r")'
# Or, clone and install locally
# git clone https://github.com/fozzieverse/fozziejoin.git
# cd fozziejoin
# Rscript.exe -e "devtools::install('./fozziejoin-r')"
Usage
Code herein is adapted from the motivating example used in the fuzzyjoin package. First, we take a list of common misspellings (and their corrected alternatives) from Wikipedia. To run in a a reasonable amount of time, we take a random sample of 1000.
library(fozziejoin)
library(tibble)
library(fuzzyjoin) # For misspellings dataset
# Load misspelling data
data(misspellings)
# Take subset of 1k records
set.seed(2016)
sub_misspellings <- misspellings[sample(nrow(misspellings), 100), ]
Next, we load a dictionary of words from the qdapDictionaries package.
library(qdapDictionaries) # For dictionary
words <- tibble::as_tibble(DICTIONARY)
Then, we run our join function.
fozzie <- fozzie_string_join(
sub_misspellings, words, method='lv',
by = c('misspelling' = 'word'), max_distance=2
)
Benchmarks
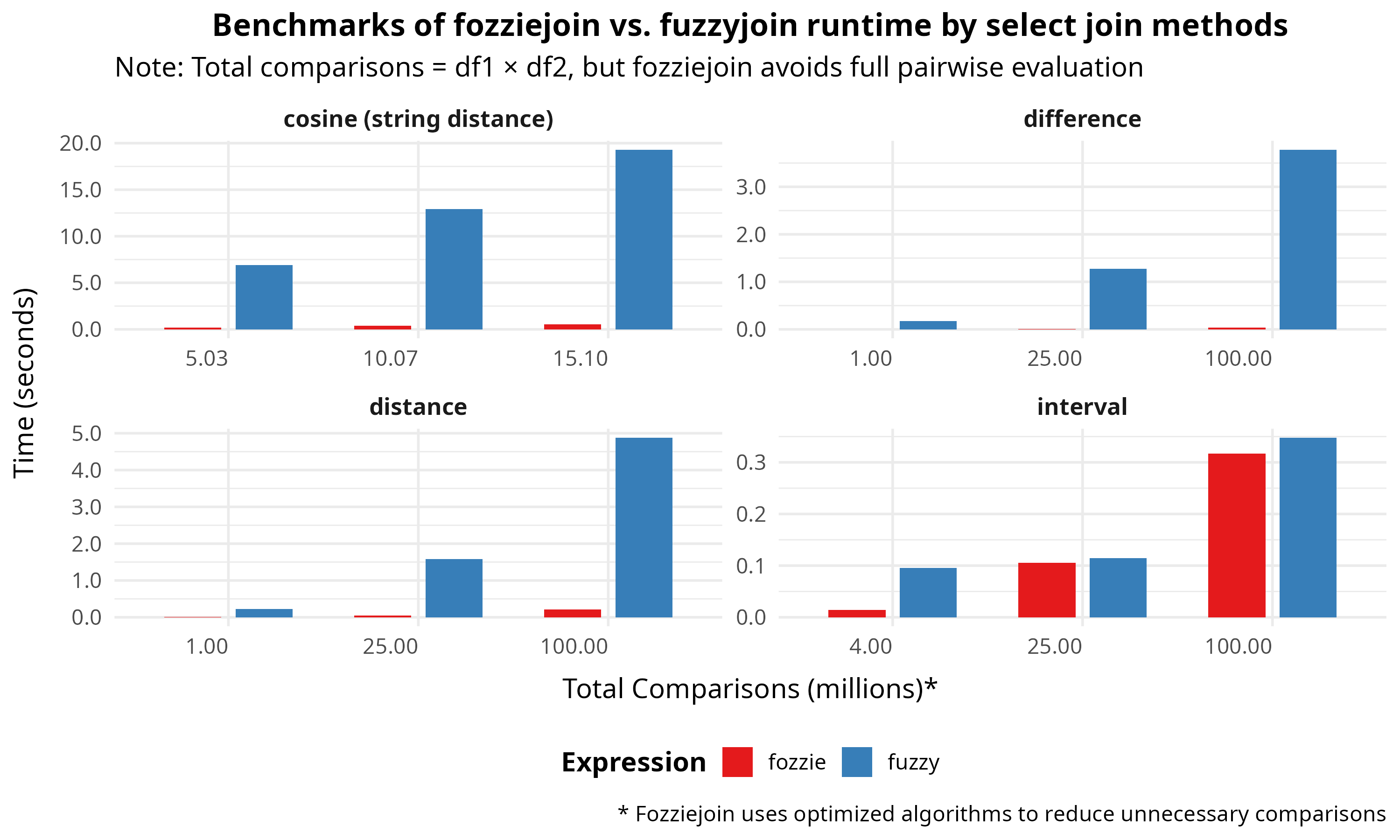
Select benchmark comparisons are below. See the benchmarks directory for the scripts ('r' subfolder) and results ('results' subfolder). For reproducibility, benchmarks are made using a GitHub workflow: see GitHub Actions Workflow for the workflow spec. Linux users will observe the largest performance gains, presumably due to the relative efficiency of parallelization via rayon.
Known behavior changes relative to fuzzyjoin
While fozziejoin is heavily inspired by fuzzyjoin, it does not seek to replicate it's behavior entirely. Please submit a GitHub issue if there are features you'd like to see! We will prioritize feature support based on community feedback.
Below are some known differences in behavior that we do not currently plan to address.
fozziejoinallowsNAvalues on the join columns specified for string distance joins.fuzzyjoinwould throw an error. This change allowsNAvalues to persist in left, right, anti, semi, and full joins. TwoNAvalues are not considered a match. We find this behavior more desirable in the case of fuzzy joins.The prefix scaling factor for Jaro-Winkler distance (
max_prefix) is an integer limiting the number of prefix characters used to boost similarity. In contrast, the analogousstringdistparameterbtis a proportion of the string length, making the prefix contribution relative rather than fixed.Some
stringdistarguments are not supported. Implementation is challenging, but not impossible. We could prioritize their inclusion if user demand were sufficient:useBytesweightuseNamesis not relevant to the final output of the fuzzy join. There is no need to implement this.
For interval joins, we allow for both
realandintegerjoin types!- The integer mode is designed to match the behavior of IRanges, which is used in
fuzzyjoin. You will need to coerce the join columns to integers to enable this mode. - The
realmode behaves more likedata.table'sfoverlaps. - An
automode (default) will determine the method to use based on the input column type
- The integer mode is designed to match the behavior of IRanges, which is used in
soundeximplementations differ slightly.- Our implementation considers multiple encodings in the case of prefixes prefixes, as is specified in the National Archives Standard.
- How consecutive similar letters and consonant separators behave is implemented differently. "Coca Cola" would match to "cuckoo" only in our system, while "overshaddowed" and "overwrought" would only match in theirs.