A Cross Between a 2D Density Plot and a Scatter Plot.
ggpointdensity
Introduces geom_pointdensity(): A cross between a scatter plot and a 2D density plot.
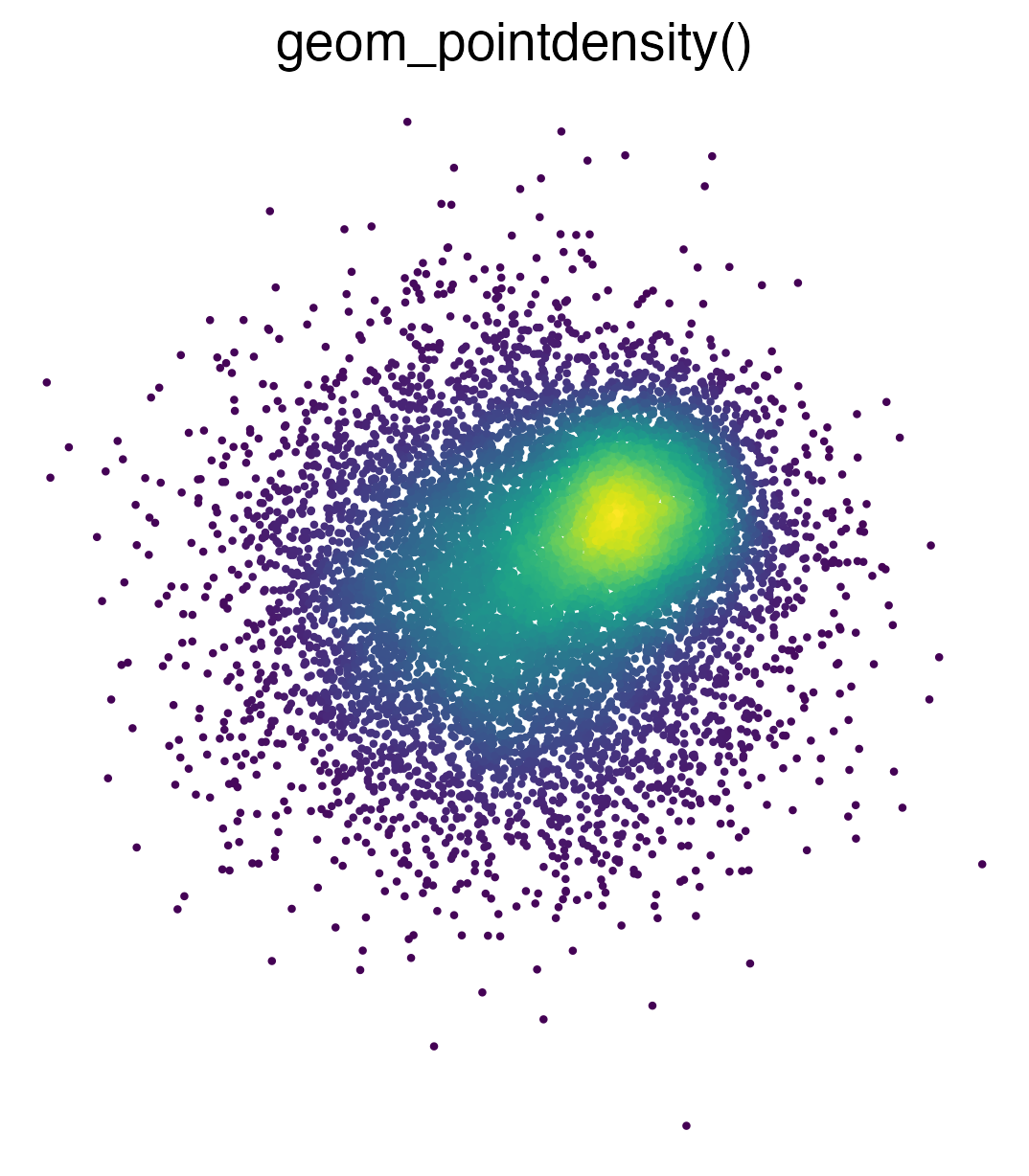
Installation
To install the package, type this command in R:
install.packages("ggpointdensity")
# Alternatively, you can install the latest
# development version from GitHub:
if (!requireNamespace("devtools", quietly = TRUE))
install.packages("devtools")
devtools::install_github("LKremer/ggpointdensity")
Motivation
There are several ways to visualize data points on a 2D coordinate system: If you have lots of data points on top of each other, geom_point() fails to give you an estimate of how many points are overlapping. geom_density2d() and geom_bin2d() solve this issue, but they make it impossible to investigate individual outlier points, which may be of interest.
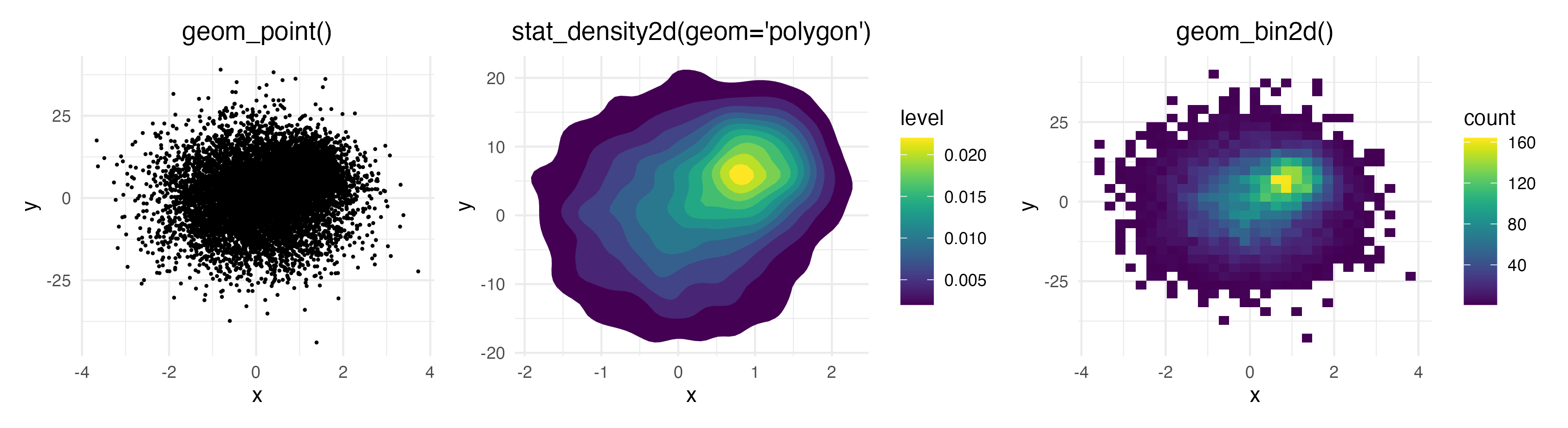
geom_pointdensity() aims to solve this problem by combining the best of both worlds: individual points are colored by the number of neighboring points. This allows you to see the overall distribution, as well as individual points.
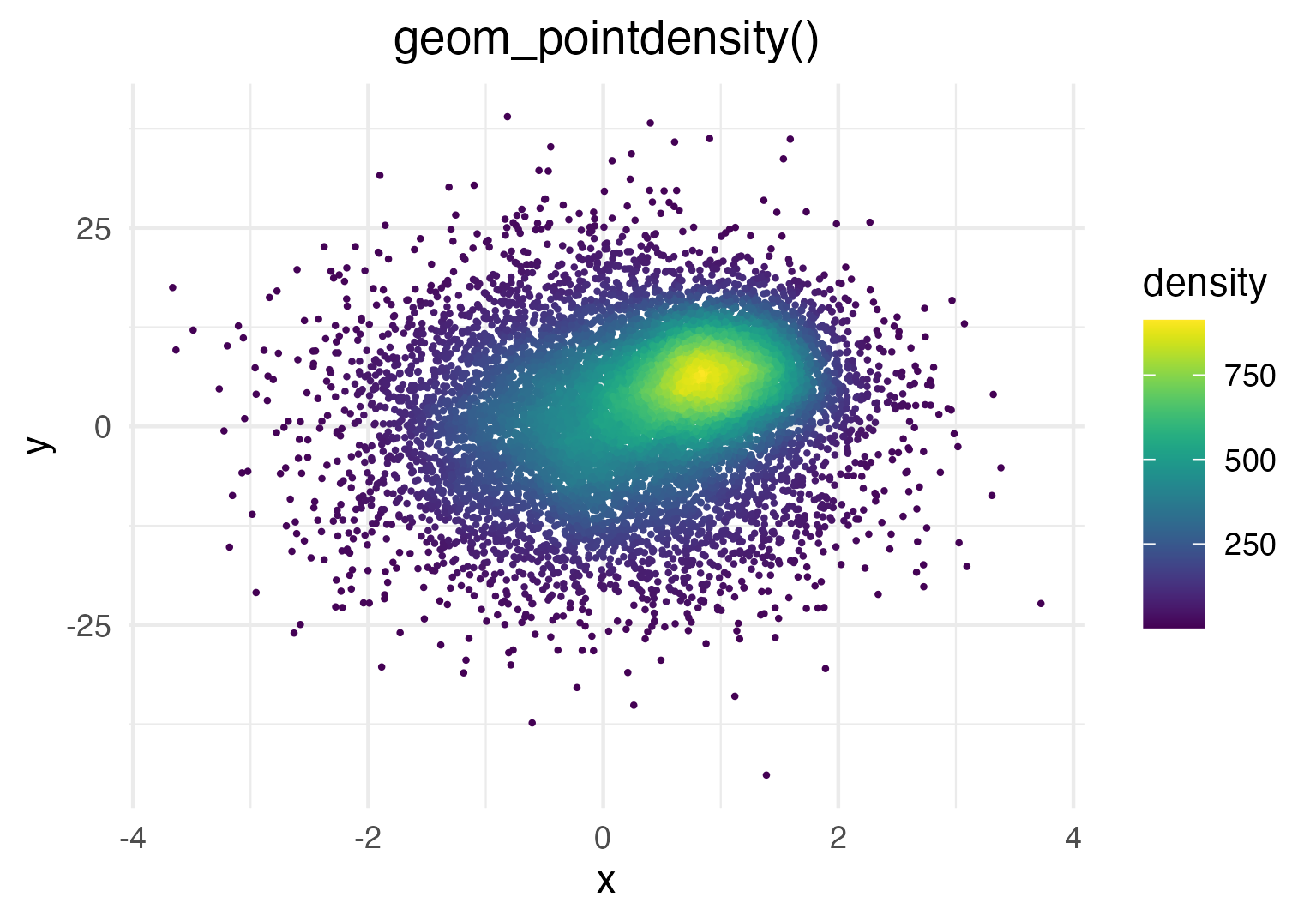
Demo
Generate some toy data and visualize it with geom_pointdensity():
library(ggplot2)
library(dplyr)
library(viridis)
library(ggpointdensity)
dat <- bind_rows(
tibble(x = rnorm(7000, sd = 1),
y = rnorm(7000, sd = 10),
group = "foo"),
tibble(x = rnorm(3000, mean = 1, sd = .5),
y = rnorm(3000, mean = 7, sd = 5),
group = "bar"))
ggplot(data = dat, mapping = aes(x = x, y = y)) +
geom_pointdensity() +
scale_color_viridis()

Each point is colored according to the number of neighboring points. The distance threshold to consider two points as neighbors (smoothing bandwidth) can be adjusted with the adjust argument, where adjust = 0.5 means use half of the default bandwidth.
ggplot(data = dat, mapping = aes(x = x, y = y)) +
geom_pointdensity(adjust = .1) +
scale_color_viridis()
ggplot(data = dat, mapping = aes(x = x, y = y)) +
geom_pointdensity(adjust = 4) +
scale_color_viridis()
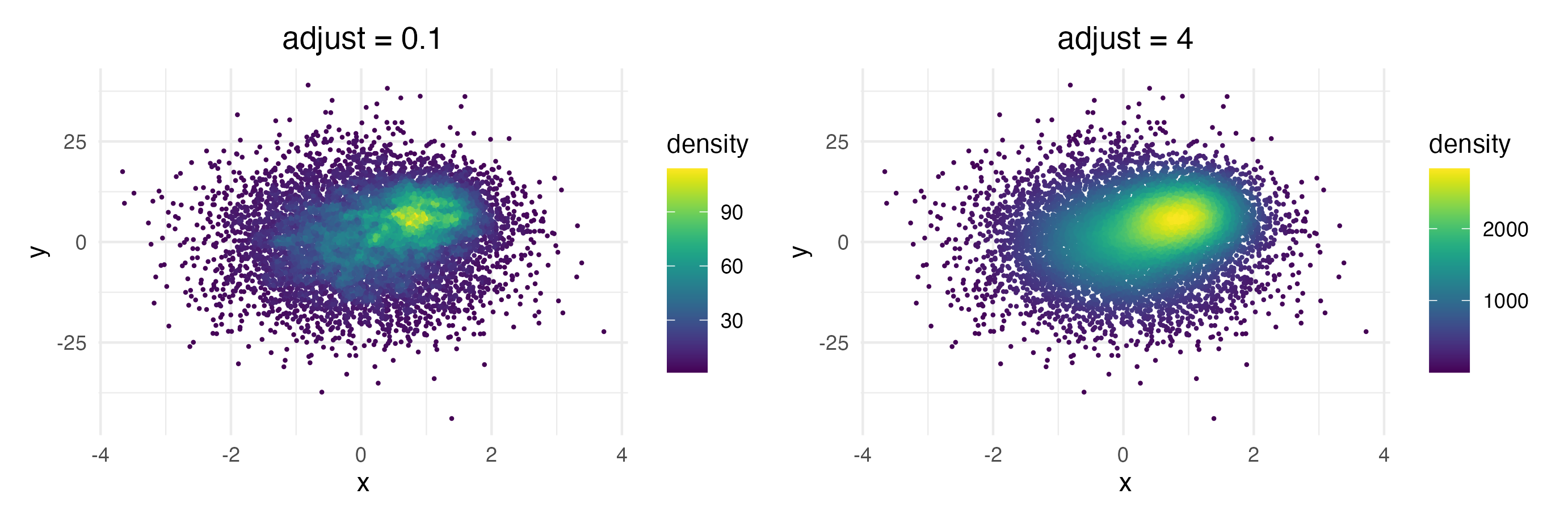
Of course you can combine the geom with standard ggplot2 features such as facets...
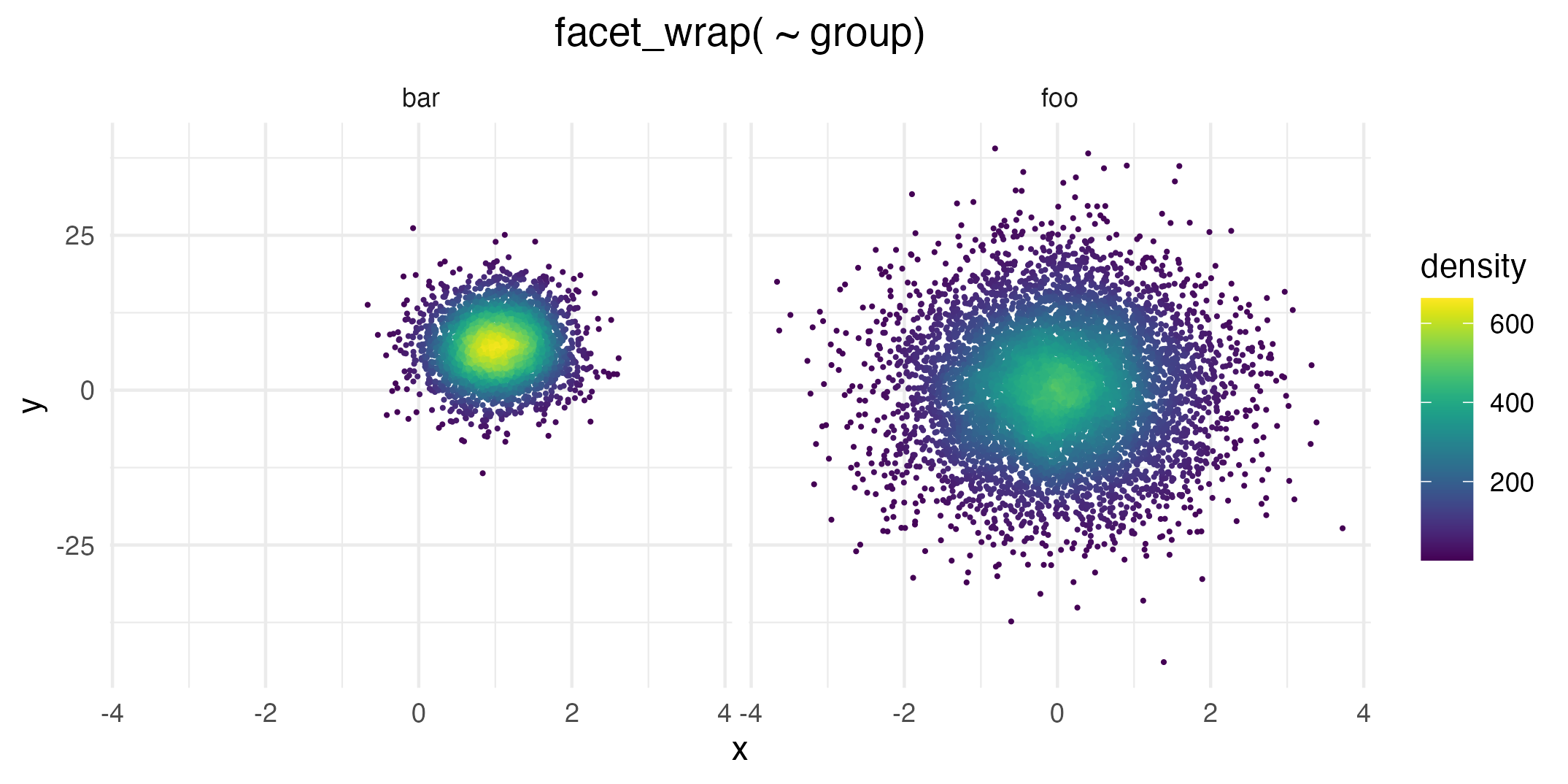
# Facetting by group
ggplot(data = dat, mapping = aes(x = x, y = y)) +
geom_pointdensity() +
scale_color_viridis() +
facet_wrap( ~ group)
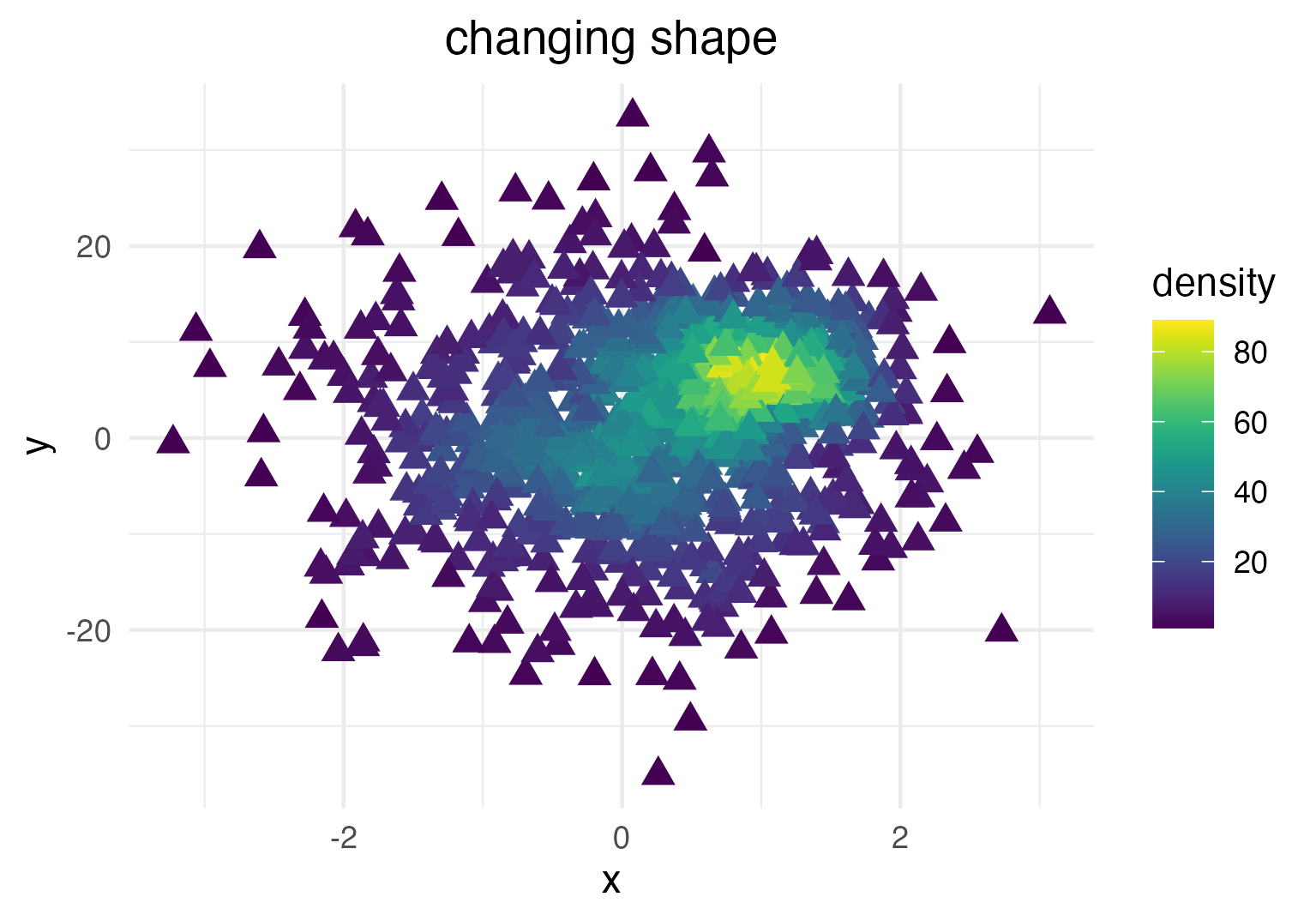
... or point shape and size:
dat_subset <- sample_frac(dat, .1) # smaller data set
ggplot(data = dat_subset, mapping = aes(x = x, y = y)) +
geom_pointdensity(size = 3, shape = 17) +
scale_color_viridis()
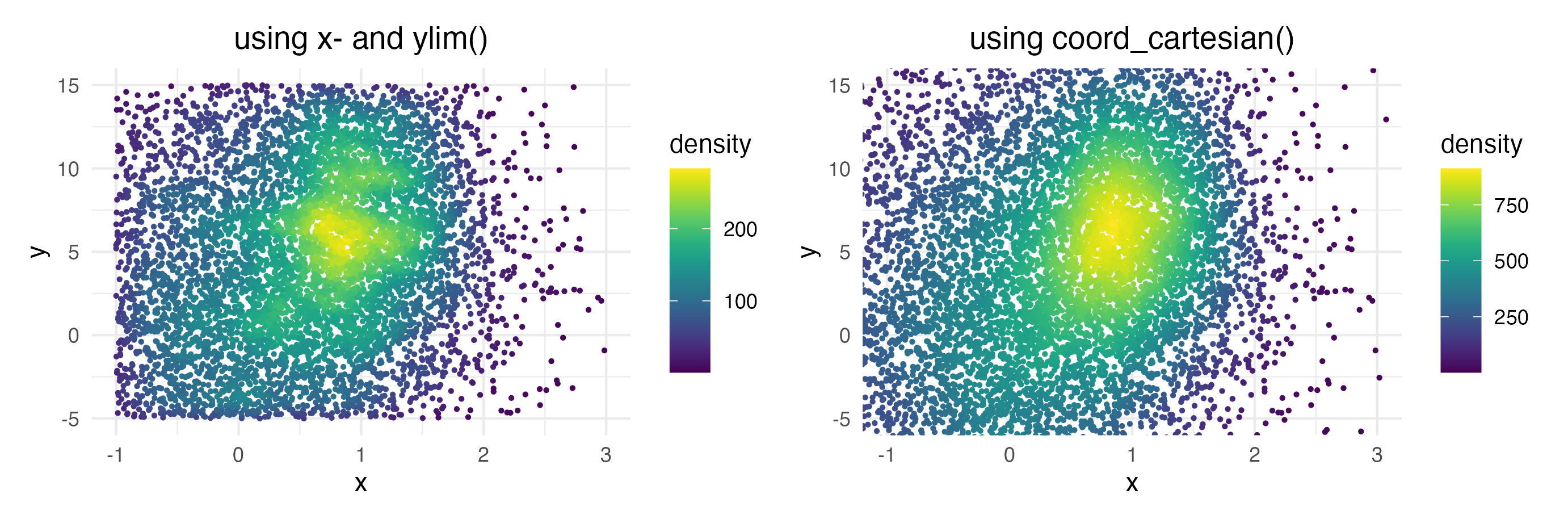
Zooming into the axis works as well, keep in mind that xlim() and ylim() change the density since they remove data points. It may be better to use coord_cartesian() instead.
ggplot(data = dat, mapping = aes(x = x, y = y)) +
geom_pointdensity() +
scale_color_viridis() +
xlim(c(-1, 3)) + ylim(c(-5, 15))
ggplot(data = dat, mapping = aes(x = x, y = y)) +
geom_pointdensity() +
scale_color_viridis() +
coord_cartesian(xlim = c(-1, 3), ylim = c(-5, 15))
Advanced usage
You can re-use or modify the density estimates using ggplot2's after_stat() function.
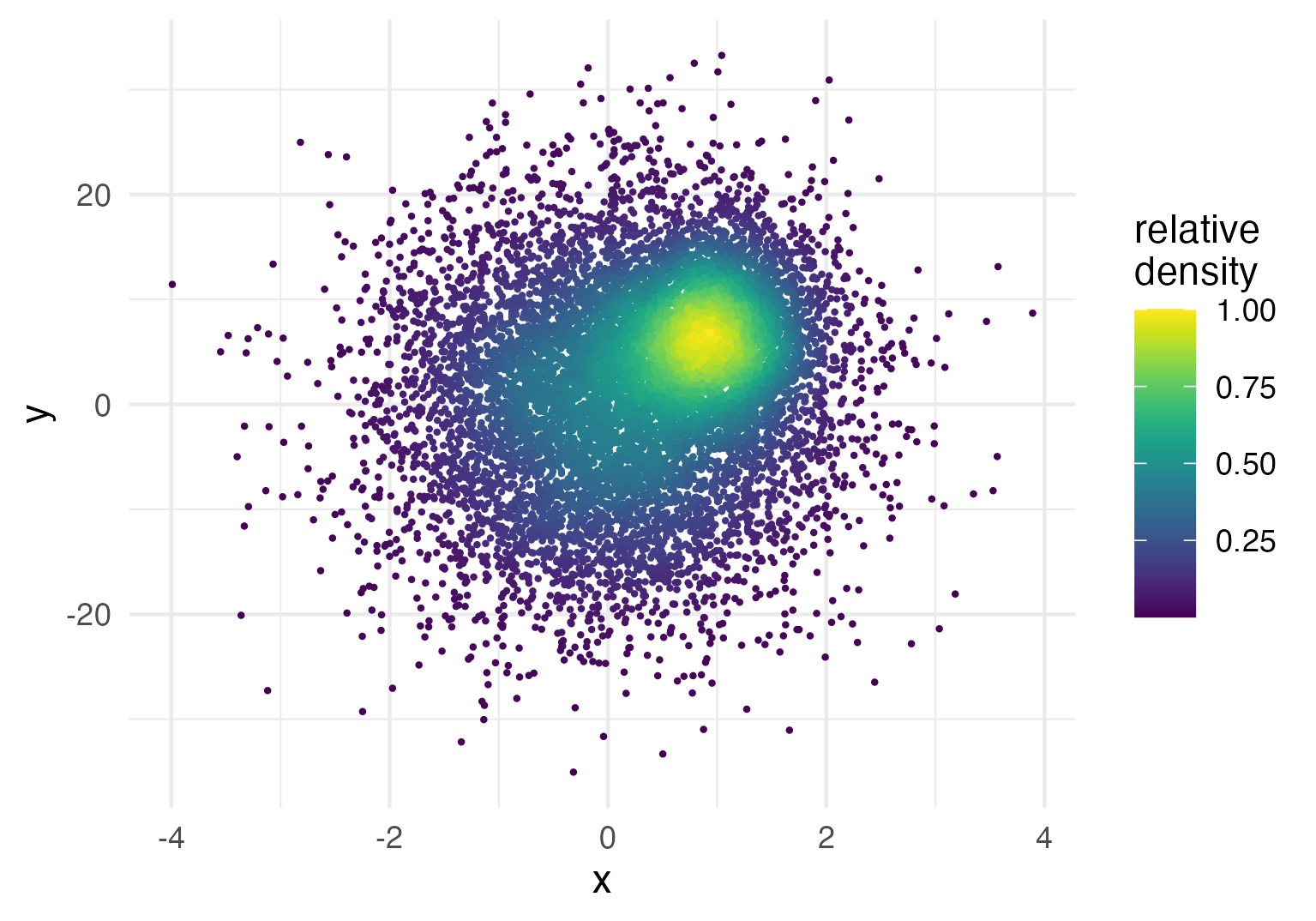
For instance, let's say you want to plot the density estimates on a relative instead of an absolute scale, i.e. scaled from 0 to 1. Of course this can be achieved by dividing the absolute density values by the maximum, but how do you access the density estimates on R code? The short answer is to use after_stat(density) inside an aesthetics mapping like so:
ggplot(data = dat,
aes(x = x, y = y, color = after_stat(density / max(density)))) +
geom_pointdensity(size = .3) +
scale_color_viridis() +
labs(color = "relative\ndensity")
For a more in-depth explanation on after_stat(), check out the relevant ggplot documentation.
Since plotting the relative density is a common use-case, we provide a little shortcut. Instead of the solution above you can simply use after_stat(ndensity). This is especially useful when facetting data, since sometimes you want to inspect the point density separately for each facet:
ggplot(data = dat,
aes( x = x, y = y, color = after_stat(ndensity))) +
geom_pointdensity( size = .25) +
scale_color_viridis() +
facet_wrap( ~ group) +
labs(color = "relative\ndensity")
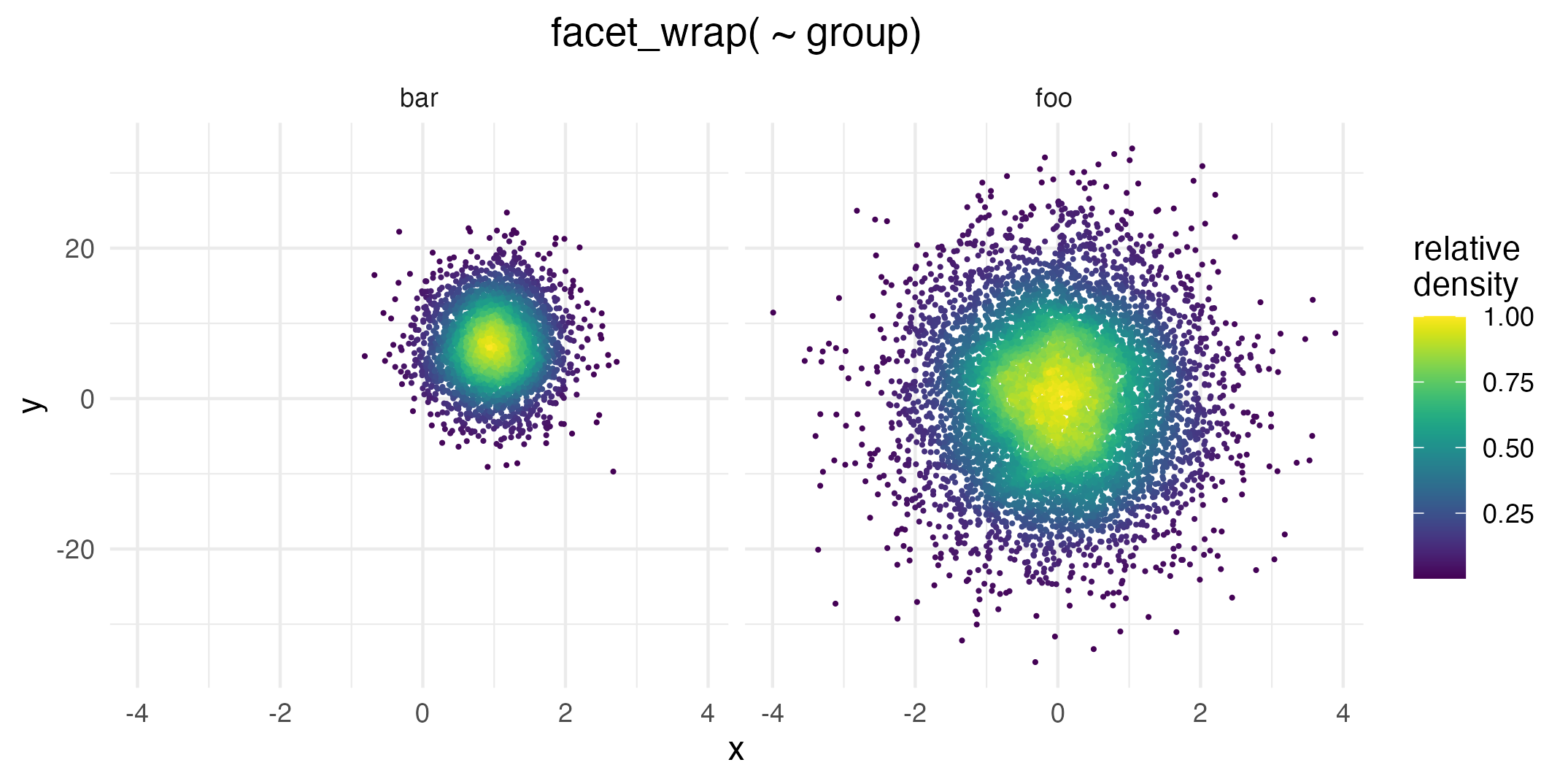
Even though the foo data group is not as dense as bar overall, this plot uses the whole color scale between 0 and 1 in both facets.
Lastly, you can use after_stat() to affect other plot aesthetics such as point size:
ggplot(data = dat,
aes(x = x, y = y, size = after_stat(1 / density ^ 1.8))) +
geom_pointdensity(adjust = .2) +
scale_color_viridis() +
scale_size_continuous(range = c(.001, 3))
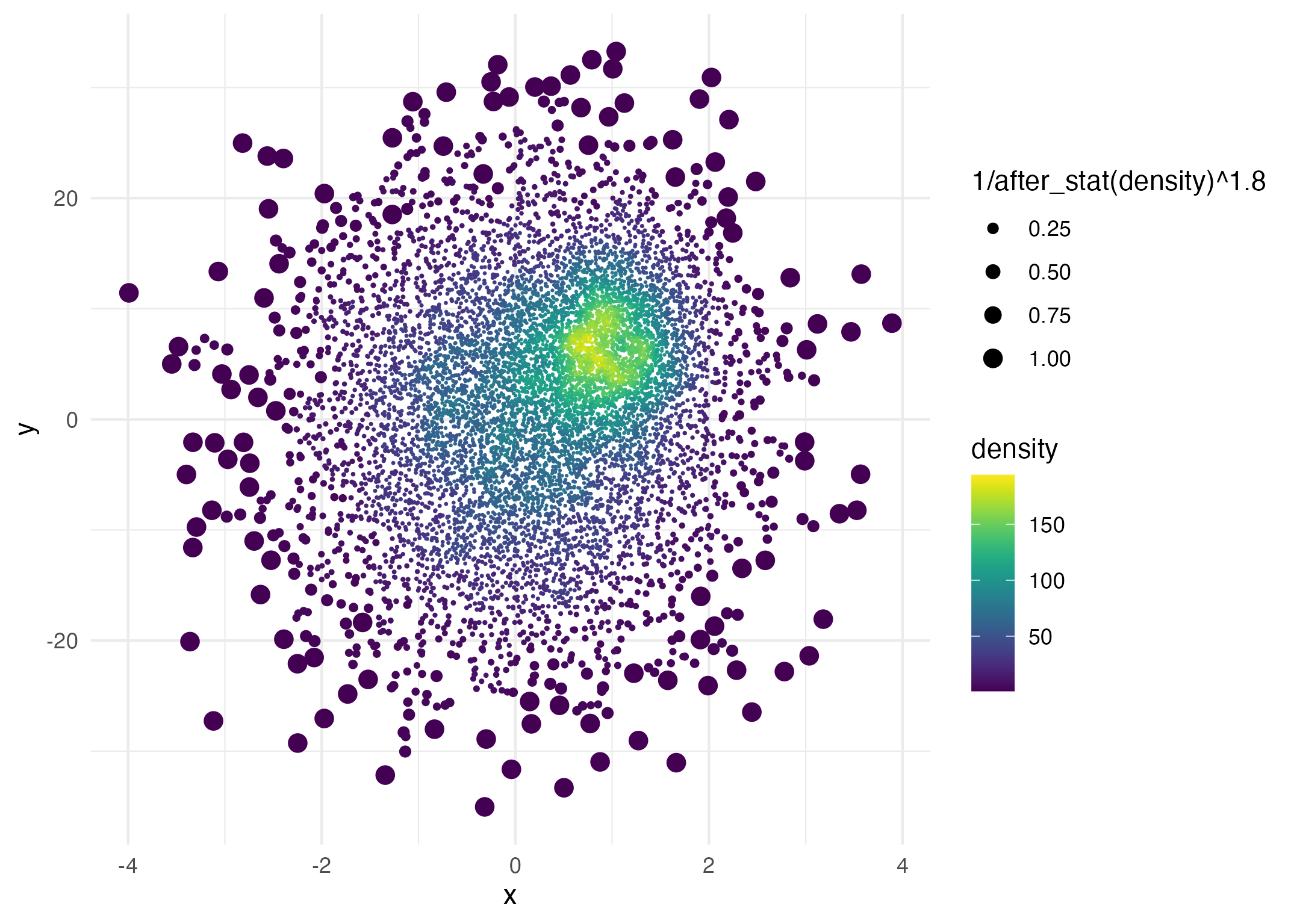
Here the point size is proportional to 1 / density ^ 1.8.
Authors
Lukas PM Kremer and Simon Anders, 2019