Higher-Level Interface of 'torch' Package to Auto-Train Neural Networks.
kindling 
Package overview
Title: Higher-Level Interface of ‘torch’ Package to Auto-Train Neural Networks
Whether you’re generating neural network architectures expressions or direct fitting/training actual models, {kindling} minimizes boilerplate code while preserving {torch}. And since this package uses {torch} as its backend, GPU/TPU devices are supported.
{kindling} also bridges the gap between {torch} and {tidymodels}. It works seamlessly with {parsnip}, {recipes}, and {workflows} to bring deep learning into your existing {tidymodels} modeling pipeline. This enables a streamlined interface for building, training, and tuning deep learning models within the familiar {tidymodels} ecosystem.
Main Features
Code generation of
{torch}expressionMultiple architectures available
- Base models interface: feedforward networks (MLP/DNN/FFNN) and recurrent variants (RNN, LSTM, GRU)
- Generalized neural network trainer that has the same topology as MLPs
Native support for titanic ML frameworks (currently supports
{tidymodels},{mlr3}for later) workflows and pipelinesFine-grained control over network depth, layer sizes, and activation functions
GPU acceleration supports via
{torch}tensors
Installation
You can install {kindling} on CRAN:
install.packages('kindling')
Or install the development version from GitHub:
# install.packages("pak")
pak::pak("joshuamarie/kindling")
## devtools::install_github("joshuamarie/kindling")
Usage: Three Levels of Interaction
{kindling} is powered by R’s metaprogramming capabilities through code generation. Generated torch::nn_module() expressions power the training functions, which in turn serve as engines for {tidymodels} integration. This architecture gives you flexibility to work at whatever abstraction level suits your task.
library(kindling)
Before starting, you need to install LibTorch, the backend of PyTorch which also the backend of {torch} R package:
torch::install_torch()
Level 1: Code Generation for torch::nn_module
At the lowest level, you can generate raw torch::nn_module code for maximum customization. Functions ending with _generator return unevaluated expressions you can inspect, modify, or execute.
Here’s how to generate a feedforward network specification:
ffnn_generator(
nn_name = "MyFFNN",
hd_neurons = c(64, 32, 16),
no_x = 10,
no_y = 1,
activations = 'relu'
)
#> torch::nn_module("MyFFNN", initialize = function ()
#> {
#> self$fc1 = torch::nn_linear(10, 64, bias = TRUE)
#> self$fc2 = torch::nn_linear(64, 32, bias = TRUE)
#> self$fc3 = torch::nn_linear(32, 16, bias = TRUE)
#> self$out = torch::nn_linear(16, 1, bias = TRUE)
#> }, forward = function (x)
#> {
#> x = self$fc1(x)
#> x = torch::nnf_relu(x)
#> x = self$fc2(x)
#> x = torch::nnf_relu(x)
#> x = self$fc3(x)
#> x = torch::nnf_relu(x)
#> x = self$out(x)
#> x
#> })
This creates a three-hidden-layer network (64 - 32 - 16 neurons) that takes 10 inputs and produces 1 output. Each hidden layer uses ReLU activation, while the output layer remains “untransformed”.
Level 2: Direct Training Interface
Skip the code generation and train models directly with your data. This approach handles all the {torch} boilerplate when training the models internally.
Let’s classify iris species:
model = ffnn(
Species ~ .,
data = iris,
hidden_neurons = c(10, 15, 7),
activations = act_funs(relu, softshrink[lambd = 0.5], elu),
loss = "cross_entropy",
epochs = 100
)
model
======================= Feedforward Neural Networks (MLP) ======================
-- FFNN Model Summary ----------------------------------------------------------
-----------------------------------------------------------------------
NN Model Type : FFNN n_predictors : 4
Number of Epochs : 100 n_response : 3
Hidden Layer Units : 10, 15, 7 reg. : None
Number of Hidden Layers : 3 Device : cpu
Pred. Type : classification :
-----------------------------------------------------------------------
-- Activation function ---------------------------------------------------------
-------------------------------------------------
1st Layer {10} : relu
2nd Layer {15} : softshrink(lambd = 0.5)
3rd Layer {7} : elu
Output Activation : No act function applied
-------------------------------------------------
For parametric activation functions like softshrink, which contains
"lambd"($\lambda$) as its parameter (the default is 1), use indexed syntax (available on v0.3.x+) e.g.softshrink[lambd = 0.5]orsoftshrink[0.5], or a string literal expression e.g."softshrink(lambd = 0.5)", to transmute the parameter value. See?kindling::act_funs()for more details.
Evaluate the prediction through predict(). The predict() method is extended for fitted models through its newdata argument.
Two kinds of predict() usage:
Without
newdatapredictions is the default to the parent data frame.predict(model) |> (\(x) table(actual = iris$Species, predicted = x))() #> predicted #> actual setosa versicolor virginica #> setosa 50 0 0 #> versicolor 0 47 3 #> virginica 0 1 49With
newdatasimply pass the new data frame as the new reference.sample_iris = dplyr::slice_sample(iris, n = 10, by = Species) predict(model, newdata = sample_iris) |> (\(x) table(actual = sample_iris$Species, predicted = x))() #> predicted #> actual setosa versicolor virginica #> setosa 10 0 0 #> versicolor 0 10 0 #> virginica 0 0 10
Level 3: Conventional tidymodels Integration
Work with neural networks just like any other {parsnip} model. This unlocks the entire {tidymodels} toolkit for preprocessing, cross-validation, and model evaluation.
# library(kindling)
# library(parsnip)
# library(yardstick)
box::use(
kindling[mlp_kindling, rnn_kindling, act_funs, args],
parsnip[fit, augment],
yardstick[metrics],
mlbench[Ionosphere] # data(Ionosphere, package = "mlbench")
)
ionosphere_data = Ionosphere[, -2]
# Train a feedforward network with parsnip
mlp_kindling(
mode = "classification",
hidden_neurons = c(128, 64),
activations = act_funs(relu, softshrink[lambd = 0.5]),
epochs = 100
) |>
fit(Class ~ ., data = ionosphere_data) |>
augment(new_data = ionosphere_data) |>
metrics(truth = Class, estimate = .pred_class)
#> # A tibble: 2 × 3
#> .metric .estimator .estimate
#> <chr> <chr> <dbl>
#> 1 accuracy binary 0.989
#> 2 kap binary 0.975
# Or try a recurrent architecture (demonstrative example with tabular data)
rnn_kindling(
mode = "classification",
hidden_neurons = c(128, 64),
activations = act_funs(relu, elu),
epochs = 100,
rnn_type = "gru"
) |>
fit(Class ~ ., data = ionosphere_data) |>
augment(new_data = ionosphere_data) |>
metrics(truth = Class, estimate = .pred_class)
#> # A tibble: 2 × 3
#> .metric .estimator .estimate
#> <chr> <chr> <dbl>
#> 1 accuracy binary 0.641
#> 2 kap binary 0
Hyperparameter Tuning & Resampling
The package has integration with {tidymodels}, so it supports hyperparameter tuning via {tune} with searchable parameters.
The current searchable parameters under {kindling}:
- Layer widths (neurons per layer)
- Network depth (number of hidden layers)
- Activation function combinations
- Output activation
- Optimizer (Type of optimization algorithm)
- Bias (choose between the presence and the absence of the bias term)
- Validation Split Proportion
- Bidirectional (boolean; only for RNN)
The searchable parameters outside from {kindling}, i.e. under {dials} package such as learn_rate() also supported.
Here’s an example:
# library(tidymodels)
box::use(
kindling[
mlp_kindling, hidden_neurons, activations, output_activation, grid_depth
],
parsnip[fit, augment],
recipes[recipe],
workflows[workflow, add_recipe, add_model],
rsample[vfold_cv],
tune[tune_grid, tune, select_best, finalize_workflow],
dials[grid_random],
yardstick[accuracy, roc_auc, metric_set, metrics]
)
mlp_tune_spec = mlp_kindling(
mode = "classification",
hidden_neurons = tune(),
activations = tune(),
output_activation = tune()
)
iris_folds = vfold_cv(iris, v = 3)
nn_wf = workflow() |>
add_recipe(recipe(Species ~ ., data = iris)) |>
add_model(mlp_tune_spec)
nn_grid_depth = grid_depth(
hidden_neurons(c(32L, 128L)),
activations(c("relu", "elu")),
output_activation(c("sigmoid", "linear")),
n_hlayer = 2,
size = 10,
type = "latin_hypercube"
)
# This is supported but limited to 1 hidden layer only
## nn_grid = grid_random(
## hidden_neurons(c(32L, 128L)),
## activations(c("relu", "elu")),
## output_activation(c("sigmoid", "linear")),
## size = 10
## )
nn_tunes = tune::tune_grid(
nn_wf,
iris_folds,
grid = nn_grid_depth
# metrics = metric_set(accuracy, roc_auc)
)
best_nn = select_best(nn_tunes)
final_nn = finalize_workflow(nn_wf, best_nn)
# Last run: 4 - 91 (relu) - 3 (sigmoid) units
final_nn_model = fit(final_nn, data = iris)
final_nn_model |>
augment(new_data = iris) |>
metrics(truth = Species, estimate = .pred_class)
#> # A tibble: 2 × 3
#> .metric .estimator .estimate
#> <chr> <chr> <dbl>
#> 1 accuracy multiclass 0.667
#> 2 kap multiclass 0.5
Resampling strategies from {rsample} will enable robust cross-validation workflows, orchestrated through the {tune} and {dials} APIs.
Variable Importance
{kindling} integrates with established variable importance methods from {NeuralNetTools} and {vip} to interpret trained neural networks. Two primary algorithms are available:
Garson’s Algorithm
garson(model, bar_plot = FALSE) #> x_names y_names rel_imp #> 1 Sepal.Width y 29.60036 #> 2 Petal.Length y 26.92774 #> 3 Petal.Width y 26.28239 #> 4 Sepal.Length y 17.18951Olden’s Algorithm
olden(model, bar_plot = FALSE) #> x_names y_names rel_imp #> 1 Petal.Width y 0.037665642 #> 2 Sepal.Length y 0.035827098 #> 3 Sepal.Width y -0.020472638 #> 4 Petal.Length y 0.009678024
Integration with {vip}
For users working within the {tidymodels} ecosystem, {kindling} models work seamlessly with the {vip} package:
box::use(
vip[vi, vip]
)
vi(model) |>
vip()
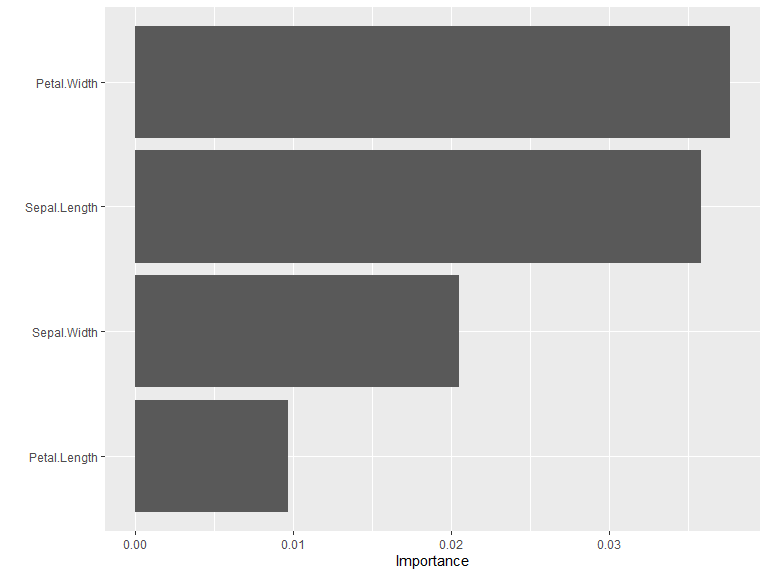
Note: Weight caching increases memory usage proportional to network size. Only enable it when you plan to compute variable importance multiple times on the same model.
References
Falbel D, Luraschi J (2023). torch: Tensors and Neural Networks with ‘GPU’ Acceleration. R package version 0.13.0, https://torch.mlverse.org, https://github.com/mlverse/torch.
Wickham H (2019). Advanced R, 2nd edition. Chapman and Hall/CRC. ISBN 978-0815384571, https://adv-r.hadley.nz/.
Goodfellow I, Bengio Y, Courville A (2016). Deep Learning. MIT Press. https://www.deeplearningbook.org/.
License
MIT + file LICENSE
Code of Conduct
Please note that the kindling project is released with a Contributor Code of Conduct. By contributing to this project, you agree to abide by its terms.