Optimal Binning Methods for Predictive Modeling and Analytics.
optbinningR
optbinningR is a native R package for optimal binning, scorecard, and monitoring workflows.
This is an independent implementation for the R community, inspired by the Python package optbinning, but rewritten for R users and R package conventions.
Install
CRAN:
install.packages("optbinningR")
CRAN status: submission is currently in process of acceptance.
GitHub:
remotes::install_github("s-rani1/optbinningR")
Local tarball:
install.packages("optbinningR_0.2.1.tar.gz", repos = NULL, type = "source")
No Python dependency for normal use
Core package usage is native R.
You do not need Python for:
- binary / multiclass / continuous optimal binning
- binning tables and plots
BinningProcess,Scorecard, monitoring, counterfactuals- 2D / piecewise / sketch / uncertainty APIs
Python parity scripts in scripts/ are optional validation utilities only.
Quick start (binary)
library(optbinningR)
d <- read.csv(system.file("extdata", "breast_cancer_mean_radius.csv", package = "optbinningR"))
y <- d$y
x <- d$x
ob <- OptimalBinning(name = "mean radius", dtype = "numerical")
ob <- fit(
ob, x, y,
algorithm = "optimal",
prebinning_method = "cart",
max_n_prebins = 20,
max_n_bins = 6,
monotonic_trend = "auto"
)
bt <- binning_table(ob)
build(bt)
plot(ob, type = "woe")
Plot previews
Binary:
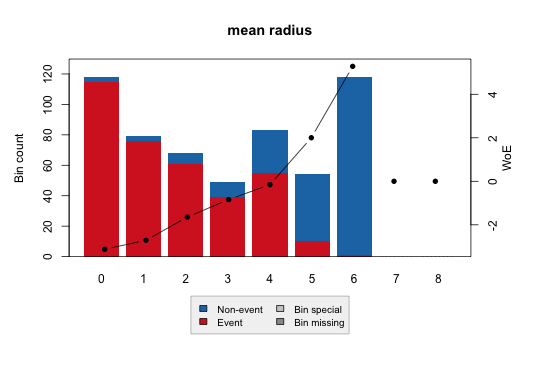
Continuous:
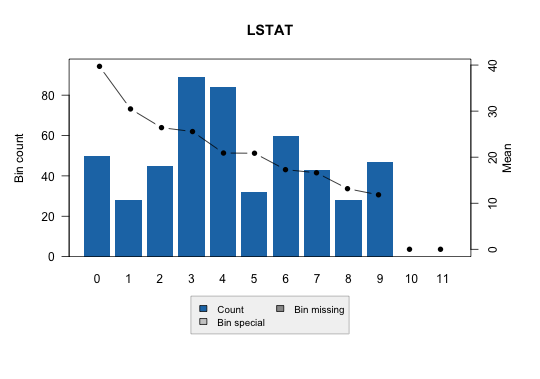
Multiclass:
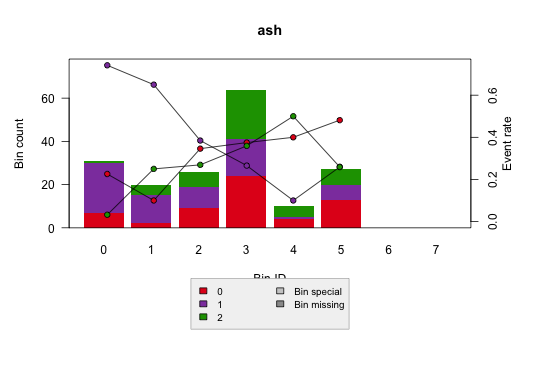
Tutorials included in this repo
These are GitHub-rendered tutorial pages (recommended for reading):
- Binary tutorial: docs/tutorials/binary.md
- Continuous tutorial: docs/tutorials/continuous.md
- Multiclass tutorial: docs/tutorials/multiclass.md
Source R Markdown files (for editing/running):
- Binary source:
inst/doc/tutorial-binary.Rmd - Continuous source:
inst/doc/tutorial-continuous.Rmd - Multiclass source:
inst/doc/tutorial-multiclass.Rmd - Getting started source:
inst/doc/getting-started.Rmd
Current scope
- Native R optimal binning: binary, multiclass, continuous targets
- Numerical and categorical variables
- Monotonic constraints:
ascending,descending,peak,valley,auto - Python-style
binning_tablehandle andbuild()output format - Plot parity work aligned to official tutorial styles
BinningProcessandScorecardworkflowsscorecard_monitoring()andcounterfactual_scorecard()- Tutorial workflow helpers:
run_fico_tutorial()andrun_telco_tutorial()
CRAN and release notes
See the CRAN release process documentation in the repository.
Contributing
Contributions are welcome via pull requests.
For substantial changes, please open an issue first to discuss scope and approach.
Issues
Bug reports and feature requests: GitHub Issues
Support
If this project is useful for your work, please consider starring the repository: s-rani1/optbinningR
Relationship to Python optbinning
- Upstream project (reference): optbinning GitHub
- Upstream docs/tutorials: optbinning documentation
optbinningR references the upstream methodology and tutorials, but the package code here is reimplemented in R for direct R usage.
Citation and attribution
If you use optbinningR, please cite:
- This repository (preferred):
GitHub “Cite this repository” page: https://github.com/s-rani1/optbinningR?tab=citation - The original
optbinningproject and references from its documentation:
https://github.com/guillermo-navas-palencia/optbinning
https://gnpalencia.org/optbinning/
Citation metadata for this repository is available from the GitHub citation tab.