Taxonomic Backbone and Name Validation Tools for Mammals of Peru.
perumammals 
Taxonomic backbone and name validation tools for the mammals of Peru.
Overview
perumammals provides a curated, standardized and programmatically accessible version of the mammal diversity of Peru as compiled by Pacheco et al. (2021): “Lista actualizada de la diversidad de los mamíferos del Perú y una propuesta para su actualización”.
This publication represents the most up-to-date and comprehensive synthesis of Peruvian mammal diversity, integrating taxonomic revisions, biogeographic information, distributional updates and the evaluation of endemic taxa.
The package includes:
- A backbone dataset of 573 mammal species known for Peru (terrestrial, arboreal, aquatic and marine).
- Taxonomic structure (Order, Family, Genus, Species, authorship).
- Endemism information.
- Ecoregional occurrence, following the scheme used by Pacheco et al. (2021).
- References and taxonomic notes as listed in the source document.
- Data structures ready for name validation, biodiversity assessments, environmental studies, species filtering, and ecoregional analyses.
The goal of the package is not to replace authoritative taxonomic databases, but to provide a stable, Peruvian-focused backbone and a set of reproducible tools that can be used in ecological, environmental, biogeographic and conservation workflows.
Context from Pacheco et al. (2021)
The backbone included in perumammals is derived directly from the annex of Pacheco et al. (2021), who synthesized decades of Peruvian mammalogy work. This list is highly relevant as it incorporates recent taxonomic updates up to November 2021, including the description of species new to science (e.g., Thomasomys antoniobracki, Oligoryzomys guille), the first Peruvian records for some bats (Eumops bonariensis), and species re-validations (e.g., Neacomys carceleni), ensuring users work with the most current classification.
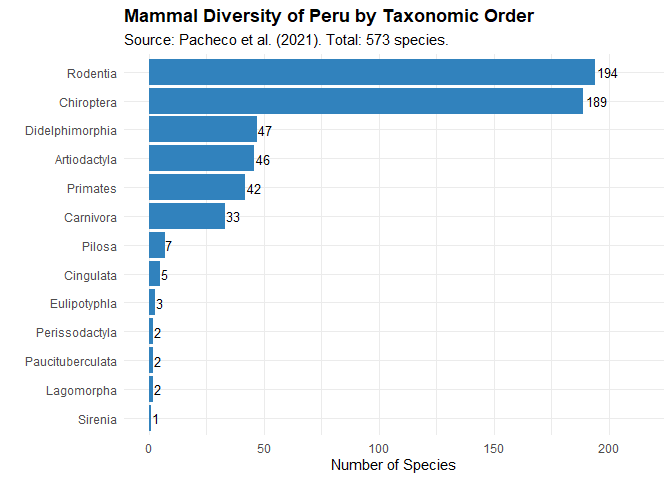
Mammalian Diversity
- 573 species documented for Peru.
- Representing 223 genera, 51 families, and 13 orders.
- Includes both terrestrial and marine mammals.
- Peru ranks among the most mammal-diverse countries worldwide.
Endemism
- 87 species are recognized as endemic to Peru, emphasizing the country’s importance for global mammalian conservation.
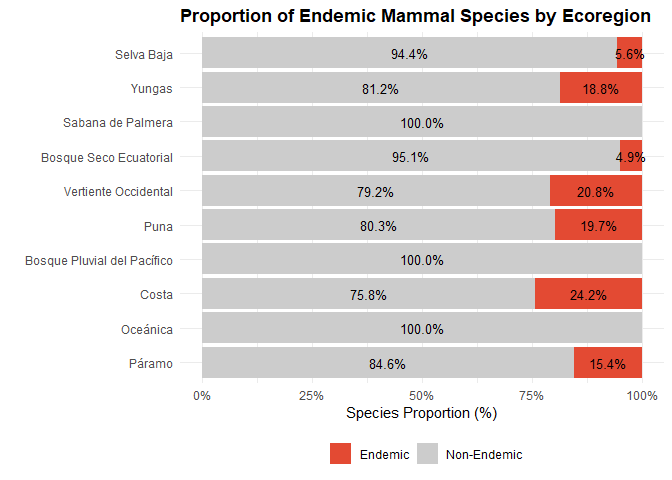
Biogeographic Ecoregions
The article assigns each species to one or more Peruvian ecoregions using the classification widely used in biogeography and conservation planning:
#> ── Peruvian Mammal Ecoregions (Brack-Egg, 1986) ────────────────────────────────
#> ℹ Number of ecoregions: 10
#> ℹ Total mammal species in Peru: 573
#>
#> Ecoregions by species richness:
#>
#> SB - Selva Baja: 320 species (55.8%)
#> YUN - Yungas: 256 species (44.7%)
#> SP - Sabana de Palmera: 83 species (14.5%)
#> BSE - Bosque Seco Ecuatorial: 81 species (14.1%)
#> VOC - Vertiente Occidental: 72 species (12.6%)
#> PUN - Puna: 71 species (12.4%)
#> BPP - Bosque Pluvial del Pacífico: 69 species (12%)
#> COS - Costa: 66 species (11.5%)
#> OCE - Oceánica: 30 species (5.2%)
#> PAR - Páramo: 26 species (4.5%)
#>
#> Use pm_by_ecoregion() to filter species by ecoregion
#> Use include_endemic = TRUE to see endemic species counts
#> ────────────────────────────────────────────────────────────────────────────────
These codes are incorporated into the package as both:
- part of the main species table, and
- a dedicated long-format table for analytical workflows.
Taxonomic Notes and Updates
Pacheco et al. (2021) incorporate:
- New species records and recent taxonomic revisions.
- Updates resulting from molecular phylogenetics and integrative taxonomy.
- Clarifications regarding problematic or doubtful taxa.
- Contextual notes on species with uncertain distributions.
What the Package Provides
Although perumammals contains functions for name validation and exploration.
The package includes four main datasets:
1. peru_mammals
A species-level backbone with:
- Taxonomy (Order, Family, Genus, Species).
- Authorship information.
- Common names when available.
- Endemic status.
- Ecoregion assignments.
- Bibliographic notes.
2. peru_mammals_ecoregions
A long-format dataset listing species–ecoregion pairs, ideal for:
- diversity summaries,
- mapping,
- ecoregional filters,
- conservation prioritization.
3. peru_mammals_ecoregions_meta
Metadata describing the ecoregion codes used across the package.
4. peru_mammals_backbone
Metadata describing:
- the source (Pacheco et al. 2021),
- year of publication,
- number of species,
- date of dataset creation during package build.
These datasets make perumammals a lightweight but powerful reference for any workflow requiring curated and Peruvian-focused mammal information.
Core Functions and Name Validation
| Category | Functionality | Conceptual Code Example |
|---|---|---|
| Name Validation | Validate species names against database | validate_peru_mammals(c("Thomasomys notatus", "Tapirus terrestris", "Unknown species")) |
| Quick Checks | Check if species occurs in Peru | is_peru_mammal("Tremarctos ornatus") |
| Endemism Query | Check endemic status | is_endemic_peru("Thomasomys notatus") |
| Match Quality | Get validation match level | match_quality_peru("Puma concolar") |
| Family Summary | List families with species counts | pm_list_families() |
| Family Filter | Filter by specific family | pm_species(family = "Cricetidae") |
| Endemic Analysis | List endemic species statistics | pm_list_endemic() |
| Endemic by Family | Filter endemics by family | pm_endemics(family = "Phyllostomidae") |
| Endemic by Ecoregion | Filter endemics by ecoregion | pm_by_ecoregion(ecoregion = "YUN", endemic = TRUE) |
Installation
## Install from CRAN (recommended)
pak::pak("perumammals")
# Development version from GitHub
# Using pak (recommended)
pak::pak("PaulESantos/perumammals")
# Or using remotes
remotes::install_github("PaulESantos/perumammals")
library(perumammals)
── Attaching perumammals ───────────────────────────────────────────────────────────────── perumammals 0.0.0.1 ──
✔ Taxonomic backbone: Pacheco et al. (2021) | Species: 573
ℹ Use pm_backbone_info() for full citation and details
Citation
If you use this package, please cite:
The package:
citation("perumammals")
#> To cite perumammals in publications, please use:
#>
#> Santos Andrade, P. E., & Gonzales Guillen, F. N. (2025). perumammals:
#> Taxonomic Backbone and Name Validation Tools for Mammals of Peru. R
#> package version 0.0.0.1. https://paulesantos.github.io/perumammals/
#>
#> The taxonomic backbone included in this package is based on:
#>
#> Pacheco, V., Cadenillas, R., Zeballos, H., Hurtado, C. M., Ruelas, D.,
#> & Pari, A. (2021). Lista actualizada de la diversidad de los mamíferos
#> del Perú y una propuesta para su actualización. Revista Peruana de
#> Biología, 28(special issue), e21019.
#> https://doi.org/10.15381/rpb.v28i4.21019
#>
#> To see these entries in BibTeX format, use 'print(<citation>,
#> bibtex=TRUE)', 'toBibtex(.)', or set
#> 'options(citation.bibtex.max=999)'.
