Description
Phylogenetic Monte Carlo.
Description
Monte Carlo based model choice for applied phylogenetics of continuous traits. Method described in Carl Boettiger, Graham Coop, Peter Ralph (2012) Is your phylogeny informative? Measuring the power of comparative methods, Evolution 66 (7) 2240-51. <doi:10.1111/j.1558-5646.2011.01574.x>.
README.md
This is a lightweight implementation of my pmc package focusing on what I think are the more common use cases (e.g. it will no longer support comparisons of a geiger model against an ouch model). Further, it does not cover many of the newer model fitting that have been implemented since pmc was first released.
The goal of this release is mostly to provide compatibility with current versions of geiger.
Getting started
Install the package:
library("devtools")
install_github("cboettig/pmc")
A trivial example with data simulated from the lambda model.
library("pmc")
library("geiger")
#> Loading required package: ape
#> Loading required package: phytools
#> Loading required package: maps
library("phytools")
phy <- sim.bdtree(n=10)
dat <- sim.char(rescale(phy, "lambda", .5), 1)[,1,]
out <- pmc(phy, dat, "BM", "lambda", nboot = 50)
#> Warning in geiger::fitContinuous(phy = tree, dat = data, model = model, :
#> Parameter estimates appear at bounds:
#> lambda
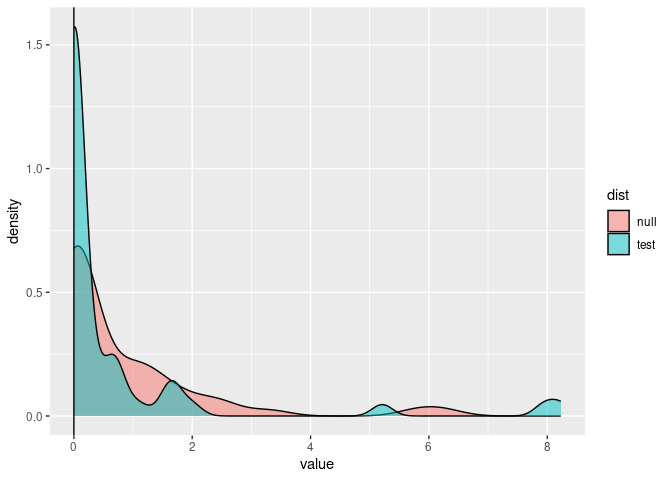
Plot the results:
dists <- data.frame(null = out$null, test = out$test)
library("ggplot2")
library("tidyr")
library("dplyr")
#>
#> Attaching package: 'dplyr'
#> The following object is masked from 'package:ape':
#>
#> where
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
dists %>%
gather(dist, value) %>%
ggplot(aes(value, fill = dist)) +
geom_density(alpha = 0.5) +
geom_vline(xintercept = out$lr)
Citation
Carl Boettiger, Graham Coop, Peter Ralph (2012) Is your phylogeny informative? Measuring the power of comparative methods, Evolution 66 (7) 2240-51. https://doi.org/10.1111/j.1558-5646.2011.01574.x.