Sample Size Analysis for Psychological Networks and More.
Sample Size Analysis for Psychological Networks
...and more
Description
powerly is an R package that implements the method by Constantin et al. (2023) for conducting sample size analysis for cross-sectional network models. The method implemented is implemented in the main function powerly(). The implementation takes the form of a three-step recursive algorithm designed to find an optimal sample size value given a model specification and an outcome measure of interest. It starts with a Monte Carlo simulation step for computing the outcome at various sample sizes. Then, it continues with a monotone curve-fitting step for interpolating the outcome. The final step employs stratified bootstrapping to quantify the uncertainty around the fitted curve.
powerly.dev
Installation
- to install from CRAN run
install.packages("powerly") - to install the latest version from GitHub run
remotes::install_github("mihaiconstantin/powerly")
Example
The code block below illustrates the main function in the package. For more information, see the documentation ?powerly, or check out the tutorials at powerly.dev.
# Suppose we want to find the sample size for observing a sensitivity of `0.6`
# with a probability of `0.8`, for a GGM true model consisting of `10` nodes
# with a density of `0.4`.
# We can run the method for an arbitrarily generated true model that matches
# those characteristics (i.e., number of nodes and density).
results <- powerly(
range_lower = 300,
range_upper = 1000,
samples = 30,
replications = 20,
measure = "sen",
statistic = "power",
measure_value = .6,
statistic_value = .8,
model = "ggm",
nodes = 10,
density = .4,
cores = 2,
verbose = TRUE
)
# Or we omit the `nodes` and `density` arguments and specify directly the edge
# weights matrix via the `model_matrix` argument.
# To get a matrix of edge weights we can use the `generate_model()` function.
true_model <- generate_model(type = "ggm", nodes = 10, density = .4)
# Then, supply the true model to the algorithm directly.
results <- powerly(
range_lower = 300,
range_upper = 1000,
samples = 30,
replications = 20,
measure = "sen",
statistic = "power",
measure_value = .6,
statistic_value = .8,
model = "ggm",
model_matrix = true_model, # Note the change.
cores = 2,
verbose = TRUE
)
To validate the results of the analysis, we can use the validate() method. For more information, see the documentation ?validate.
# Validate the recommendation obtained during the analysis.
validation <- validate(results, cores = 2)
To visualize the results, we can use the plot function and indicate the step that should be plotted.
# Step 1.
plot(results, step = 1)
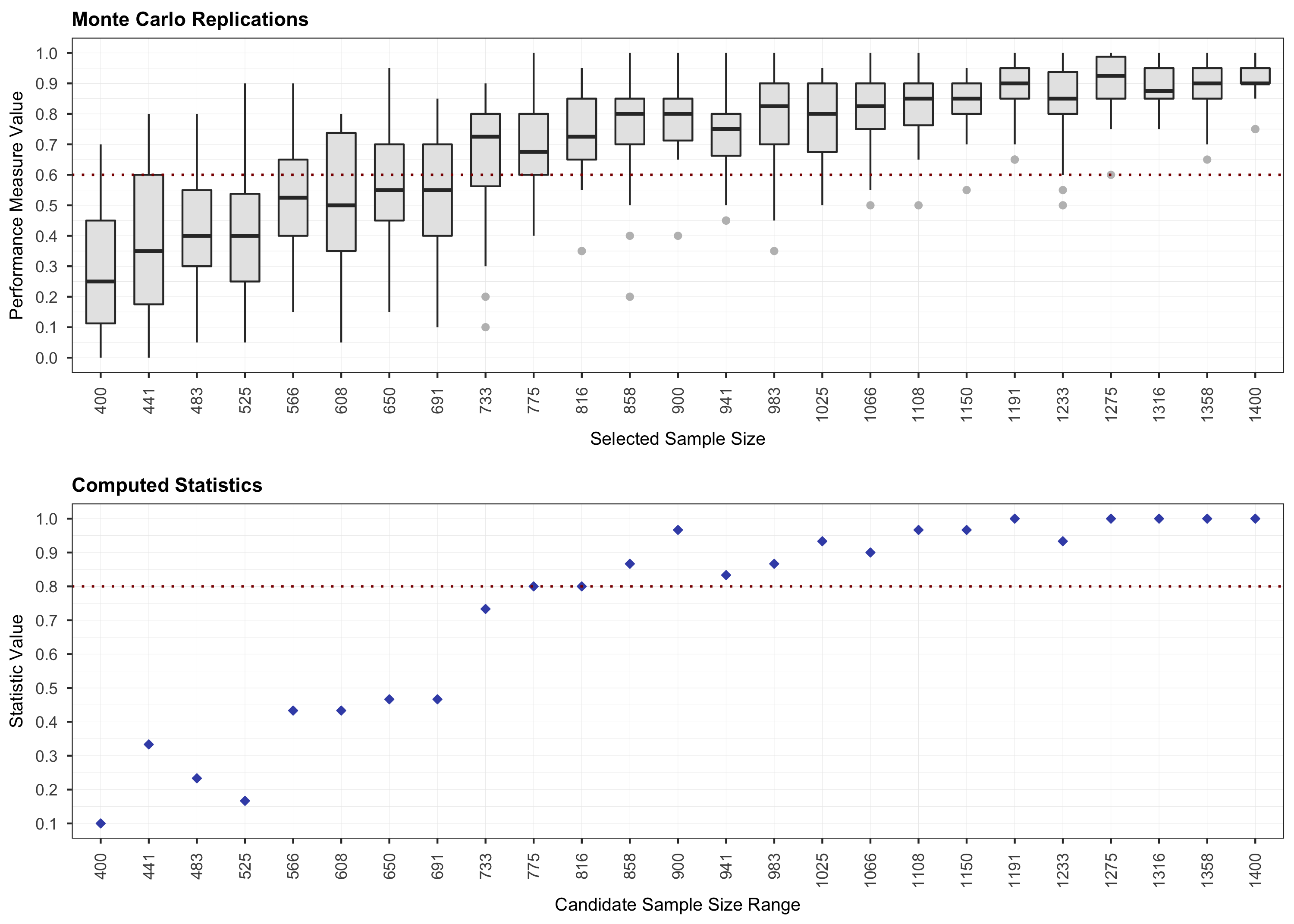
# Step 2.
plot(results, step = 2)
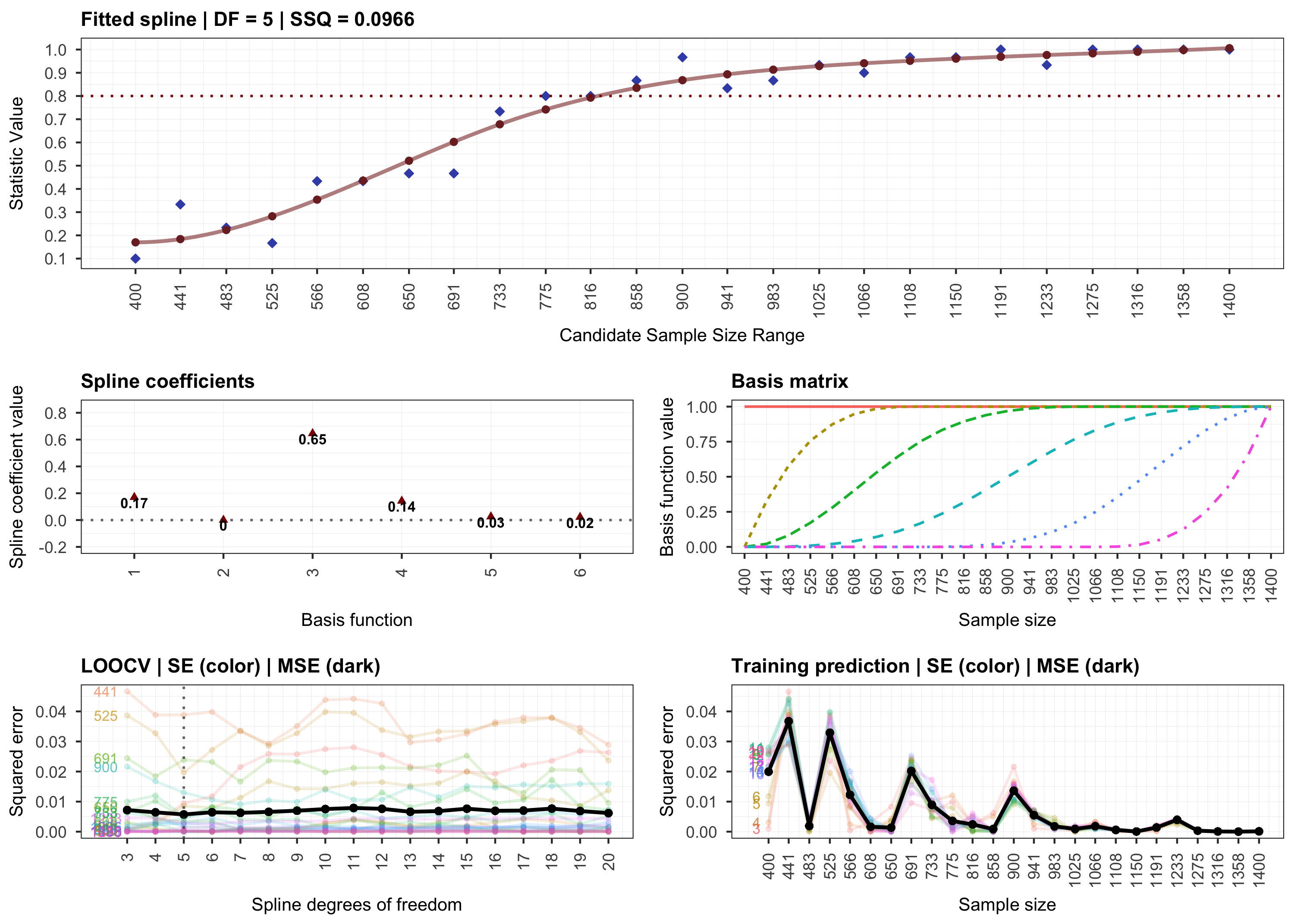
# Step 3.
plot(results, step = 3)
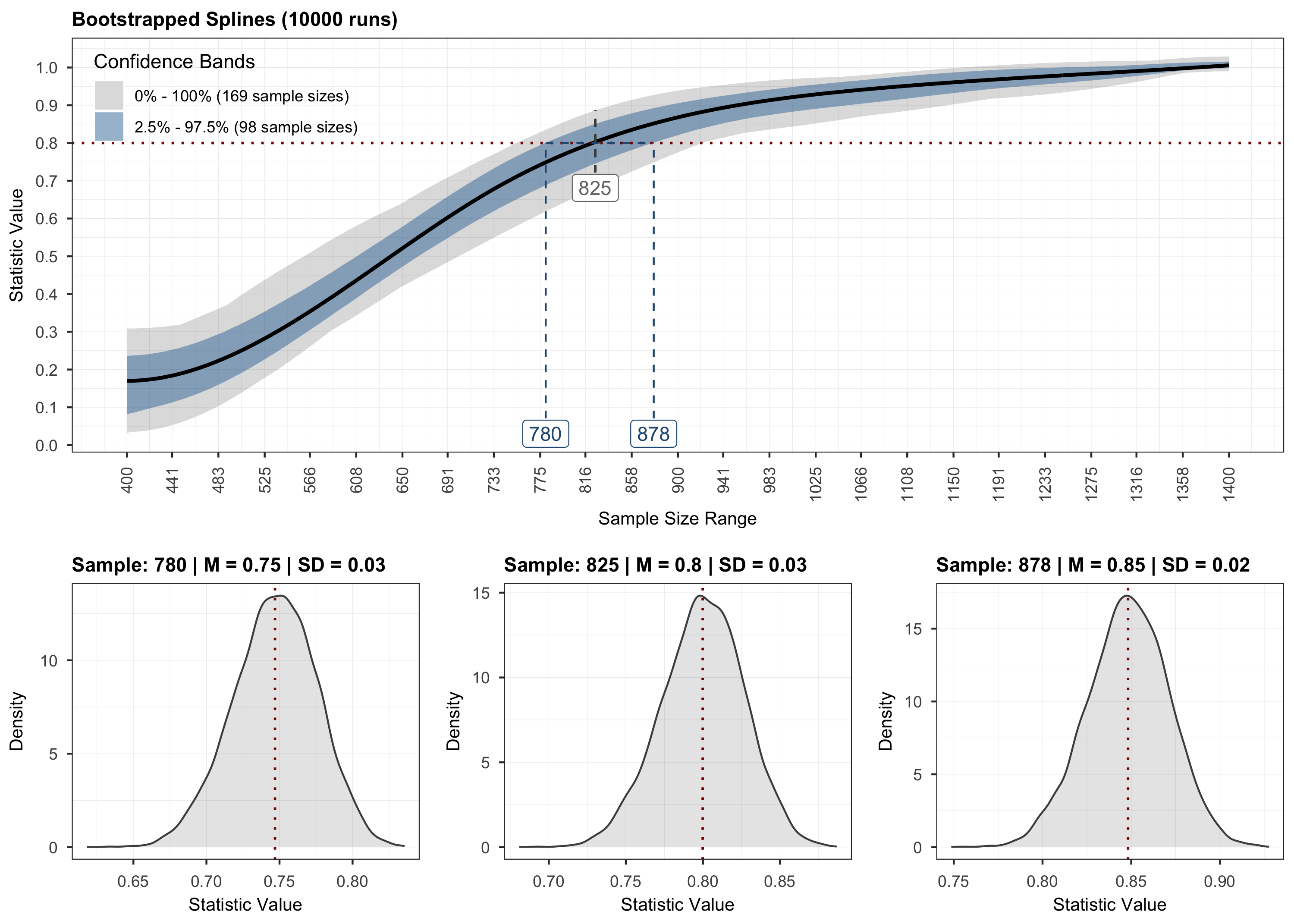
# Validation.
plot(validation)
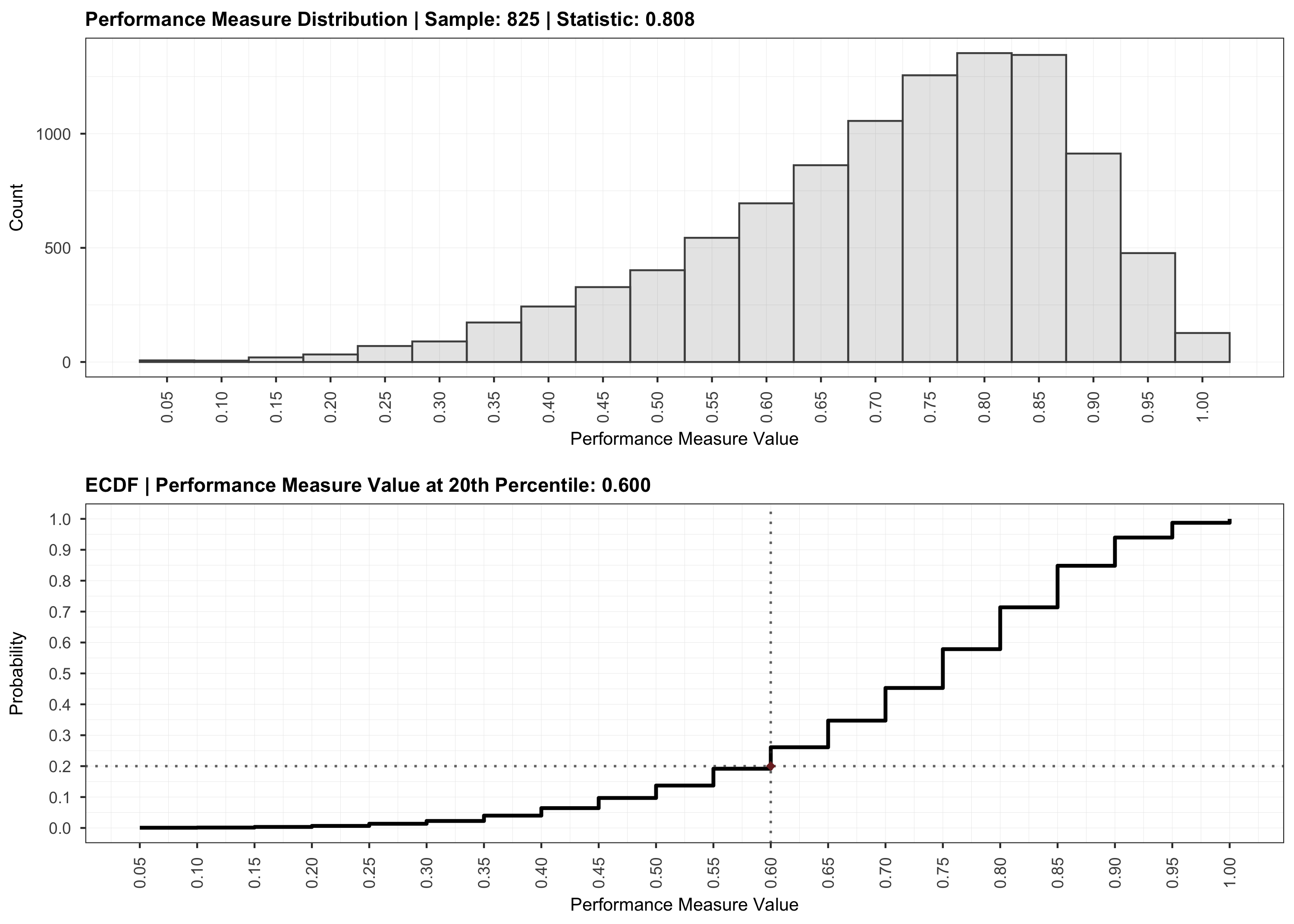
Contributing
- To support a new model, performance measure, or statistic, please open a pull request on GitHub.
- To request a new model, performance measure, or statistic, please open an issue on GitHub. If possible, also include references discussing the topics you are requesting.
Poster
License
The code in this repository is licensed under the MIT license.
To use powerly please cite:
- Constantin, M. A., Schuurman, N. K., & Vermunt, J. K. (2023). A General Monte Carlo Method for Sample Size Analysis in the Context of Network Models. Psychological Methods. https://doi.org/10.1037/met0000555