Compiler for R.
quickr 
The goal of quickr is to make your R code run quicker.
Overview
R is an extremely flexible and dynamic language, but that flexibility and dynamicism can come at the expense of speed. This package lets you trade back some of that flexibility for some speed, for the context of a single function.
The main exported function is quick(), here is how you use it.
library(quickr)
convolve <- quick(function(a, b) {
declare(type(a = double(NA)),
type(b = double(NA)))
ab <- double(length(a) + length(b) - 1)
for (i in seq_along(a)) {
for (j in seq_along(b)) {
ab[i+j-1] <- ab[i+j-1] + a[i] * b[j]
}
}
ab
})
quick() returns a quicker R function. How much quicker? Let’s benchmark it! For reference, we’ll also compare it to a pure-C implementation.
slow_convolve <- function(a, b) {
declare(type(a = double(NA)),
type(b = double(NA)))
ab <- double(length(a) + length(b) - 1)
for (i in seq_along(a)) {
for (j in seq_along(b)) {
ab[i+j-1] <- ab[i+j-1] + a[i] * b[j]
}
}
ab
}
library(quickr)
quick_convolve <- quick(slow_convolve)
convolve_c <- inline::cfunction(
sig = c(a = "SEXP", b = "SEXP"), body = r"({
int na, nb, nab;
double *xa, *xb, *xab;
SEXP ab;
a = PROTECT(Rf_coerceVector(a, REALSXP));
b = PROTECT(Rf_coerceVector(b, REALSXP));
na = Rf_length(a); nb = Rf_length(b); nab = na + nb - 1;
ab = PROTECT(Rf_allocVector(REALSXP, nab));
xa = REAL(a); xb = REAL(b); xab = REAL(ab);
for(int i = 0; i < nab; i++) xab[i] = 0.0;
for(int i = 0; i < na; i++)
for(int j = 0; j < nb; j++)
xab[i + j] += xa[i] * xb[j];
UNPROTECT(3);
return ab;
})")
a <- runif (100000); b <- runif (100)
timings <- bench::mark(
r = slow_convolve(a, b),
quickr = quick_convolve(a, b),
c = convolve_c(a, b),
min_time = 2
)
timings
#> # A tibble: 3 × 6
#> expression min median `itr/sec` mem_alloc `gc/sec`
#> <bch:expr> <bch:tm> <bch:tm> <dbl> <bch:byt> <dbl>
#> 1 r 1.05s 1.05s 0.955 847KB 0.955
#> 2 quickr 4.8ms 5.09ms 195. 782KB 3.07
#> 3 c 4.73ms 5.04ms 196. 782KB 3.09
plot(timings) + bench::scale_x_bench_time(base = NULL)
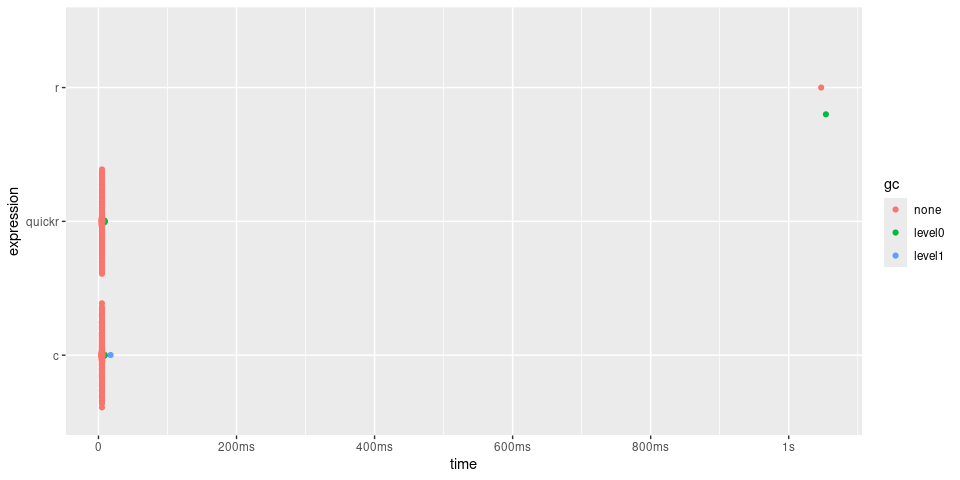
In the case of convolve(), quick() returns a function approximately 200 times quicker, giving similar performance to the C function.
quick() can accelerate any R function, with some restrictions:
- Function arguments must have their types and shapes declared using
declare(). - Only atomic vectors, matrices, and array are currently supported:
integer,double,logical, andcomplex. - The return value must be an atomic array (e.g., not a list)
- Named variables must have consistent shapes throughout their lifetimes.
NAvalues are not supported.- Only a subset of R’s vocabulary is currently supported.
#> [1] - : != ( [ [<- {
#> [8] * / & && %/% %% ^
#> [15] + < <- <= = == >
#> [22] >= | || Arg Conj Fortran Im
#> [29] Mod Re abs acos asin atan c
#> [36] cat cbind ceiling character cos declare double
#> [43] exp floor for if ifelse integer length
#> [50] log log10 logical matrix max min numeric
#> [57] print prod raw seq sin sqrt sum
#> [64] tan which.max which.min
Many of these restrictions are expected to be relaxed as the project matures. However, quickr is intended primarily for high-performance numerical computing, so features like polymorphic dispatch or support for complex or dynamic types are out of scope.
declare(type()) syntax:
The shape and mode of all function arguments must be declared. Local and return variables may optionally also be declared.
declare(type()) also has support for declaring size constraints, or size relationships between variables. Here are some examples of declare calls:
declare(type(x = double(NA))) # x is a 1-d double vector of any length
declare(type(x = double(10))) # x is a 1-d double vector of length 10
declare(type(x = double(1))) # x is a scalar double
declare(type(x = integer(2, 3))) # x is a 2-d integer matrix with dim (2, 3)
declare(type(x = integer(NA, 3))) # x is a 2-d integer matrix with dim (<any>, 3)
# x is a 4-d logical matrix with dim (<any>, 24, 24, 3)
declare(type(x = logical(NA, 24, 24, 3)))
# x and y are 1-d double vectors of any length
declare(type(x = double(NA)),
type(y = double(NA)))
# x and y are 1-d double vectors of the same length
declare(
type(x = double(n)),
type(y = double(n)),
)
# x and y are 1-d double vectors, where length(y) == length(x) + 2
declare(type(x = double(n)),
type(y = double(n+2)))
More examples:
viterbi
The Viterbi algorithm is an example of a dynamic programming algorithm within the family of Hidden Markov Models (https://en.wikipedia.org/wiki/Viterbi_algorithm). Here, quick() makes the viterbi() approximately 50 times faster.
slow_viterbi <- function(observations, states, initial_probs, transition_probs, emission_probs) {
declare(
type(observations = integer(num_steps)),
type(states = integer(num_states)),
type(initial_probs = double(num_states)),
type(transition_probs = double(num_states, num_states)),
type(emission_probs = double(num_states, num_obs)),
)
trellis <- matrix(0, nrow = length(states), ncol = length(observations))
backpointer <- matrix(0L, nrow = length(states), ncol = length(observations))
trellis[, 1] <- initial_probs * emission_probs[, observations[1]]
for (step in 2:length(observations)) {
for (current_state in 1:length(states)) {
probabilities <- trellis[, step - 1] * transition_probs[, current_state]
trellis[current_state, step] <- max(probabilities) * emission_probs[current_state, observations[step]]
backpointer[current_state, step] <- which.max(probabilities)
}
}
path <- integer(length(observations))
path[length(observations)] <- which.max(trellis[, length(observations)])
for (step in seq(length(observations) - 1, 1)) {
path[step] <- backpointer[path[step + 1], step + 1]
}
out <- states[path]
out
}
quick_viterbi <- quick(slow_viterbi)
set.seed(1234)
num_steps <- 16
num_states <- 8
num_obs <- 16
observations <- sample(1:num_obs, num_steps, replace = TRUE)
states <- 1:num_states
initial_probs <- runif (num_states)
initial_probs <- initial_probs / sum(initial_probs) # normalize to sum to 1
transition_probs <- matrix(runif (num_states * num_states), nrow = num_states)
transition_probs <- transition_probs / rowSums(transition_probs) # normalize rows
emission_probs <- matrix(runif (num_states * num_obs), nrow = num_states)
emission_probs <- emission_probs / rowSums(emission_probs) # normalize rows
timings <- bench::mark(
slow_viterbi = slow_viterbi(observations, states, initial_probs,
transition_probs, emission_probs),
quick_viterbi = quick_viterbi(observations, states, initial_probs,
transition_probs, emission_probs)
)
timings
#> # A tibble: 2 × 6
#> expression min median `itr/sec` mem_alloc `gc/sec`
#> <bch:expr> <bch:tm> <bch:tm> <dbl> <bch:byt> <dbl>
#> 1 slow_viterbi 143.04µs 162µs 6047. 178KB 14.5
#> 2 quick_viterbi 2.43µs 2.53µs 368609. 0B 0
plot(timings)
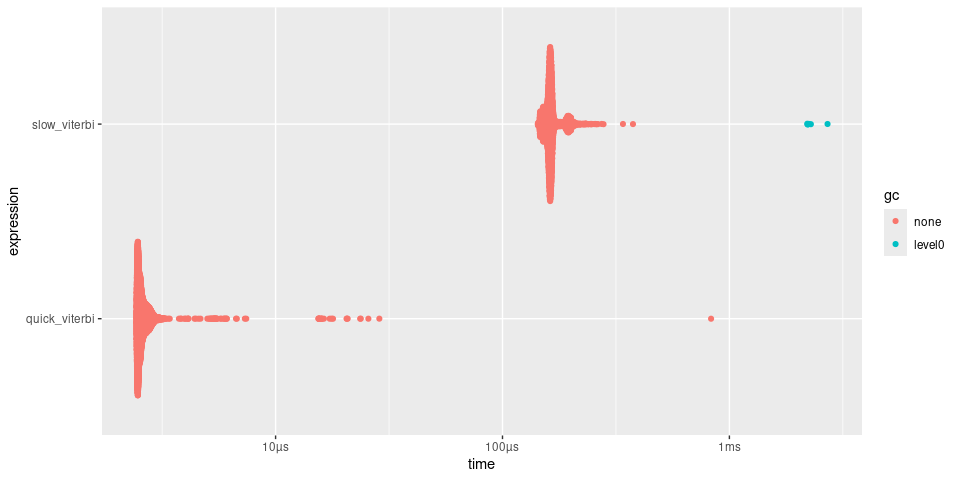
Diffusion simulation
Simulate how heat spreads over time across a 2D grid, using the finite difference method applied to the Heat Equation.
Here, quick() returns a function over 100 time faster.
diffuse_heat <- function(nx, ny, dx, dy, dt, k, steps) {
declare(
type(nx = integer(1)),
type(ny = integer(1)),
type(dx = integer(1)),
type(dy = integer(1)),
type(dt = double(1)),
type(k = double(1)),
type(steps = integer(1))
)
# Initialize temperature grid
temp <- matrix(0, nx, ny)
temp[nx / 2, ny / 2] <- 100 # Initial heat source in the center
# Time stepping
for (step in seq_len(steps)) {
# Apply boundary conditions
temp[1, ] <- 0
temp[nx, ] <- 0
temp[, 1] <- 0
temp[, ny] <- 0
# Update using finite differences
temp_new <- temp
for (i in 2:(nx - 1)) {
for (j in 2:(ny - 1)) {
temp_new[i, j] <- temp[i, j] + k * dt *
((temp[i + 1, j] - 2 * temp[i, j] + temp[i - 1, j]) /
dx ^ 2 + (temp[i, j + 1] - 2 * temp[i, j] + temp[i, j - 1]) / dy ^ 2)
}
}
temp <- temp_new
}
temp
}
quick_diffuse_heat <- quick(diffuse_heat)
# Parameters
nx <- 100L # Grid size in x
ny <- 100L # Grid size in y
dx <- 1L # Grid spacing
dy <- 1L # Grid spacing
dt <- 0.01 # Time step
k <- 0.1 # Thermal diffusivity
steps <- 500L # Number of time steps
timings <- bench::mark(
diffuse_heat = diffuse_heat(nx, ny, dx, dy, dt, k, steps),
quick_diffuse_heat = quick_diffuse_heat(nx, ny, dx, dy, dt, k, steps)
)
#> Warning: Some expressions had a GC in every iteration; so filtering is
#> disabled.
summary(timings, relative = TRUE)
#> Warning: Some expressions had a GC in every iteration; so filtering is
#> disabled.
#> # A tibble: 2 × 6
#> expression min median `itr/sec` mem_alloc `gc/sec`
#> <bch:expr> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 diffuse_heat 127. 122. 1 1014. Inf
#> 2 quick_diffuse_heat 1 1 121. 1 NaN
plot(timings)
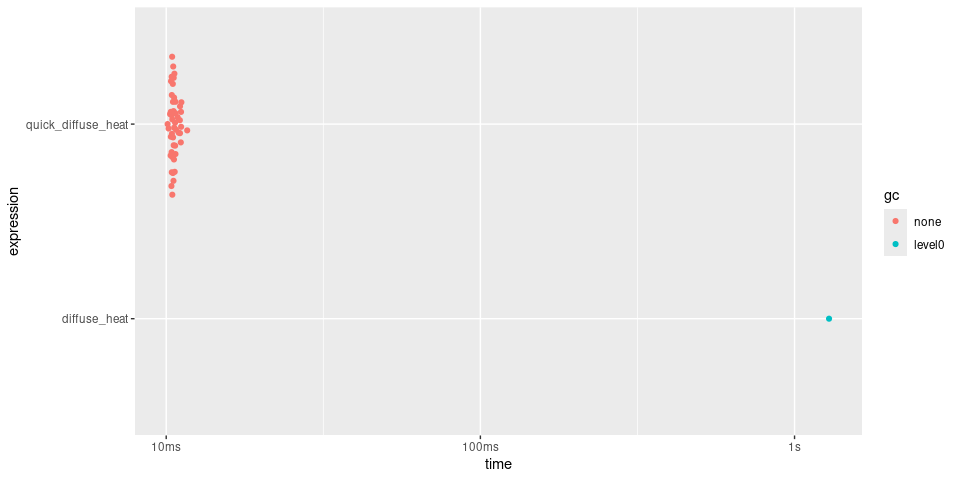
Rolling Mean
Here is quickr used to calculate a rolling mean. Note that the CRAN package RcppRoll already provides a highly optimized rolling mean, which we include in the benchmarks for comparison.
slow_roll_mean <- function(x, weights, normalize = TRUE) {
declare(
type(x = double(NA)),
type(weights = double(NA)),
type(normalize = logical(1))
)
out <- double(length(x) - length(weights) + 1)
n <- length(weights)
if (normalize)
weights <- weights/sum(weights)*length(weights)
for(i in seq_along(out)) {
out[i] <- sum(x[i:(i+n-1)] * weights) / length(weights)
}
out
}
quick_roll_mean <- quick(slow_roll_mean)
x <- dnorm(seq(-3, 3, len = 100000))
weights <- dnorm(seq(-1, 1, len = 100))
timings <- bench::mark(
r = slow_roll_mean(x, weights),
rcpp = RcppRoll::roll_mean(x, weights = weights),
quickr = quick_roll_mean(x, weights = weights)
)
#> Warning: Some expressions had a GC in every iteration; so filtering is
#> disabled.
timings
#> # A tibble: 3 × 6
#> expression min median `itr/sec` mem_alloc `gc/sec`
#> <bch:expr> <bch:tm> <bch:tm> <dbl> <bch:byt> <dbl>
#> 1 r 105.4ms 109.13ms 7.42 124.31MB 22.3
#> 2 rcpp 19.7ms 19.84ms 49.4 4.44MB 1.98
#> 3 quickr 7ms 7.09ms 138. 781.35KB 2.00
timings$expression <- factor(names(timings$expression), rev(names(timings$expression)))
plot(timings) + bench::scale_x_bench_time(base = NULL)
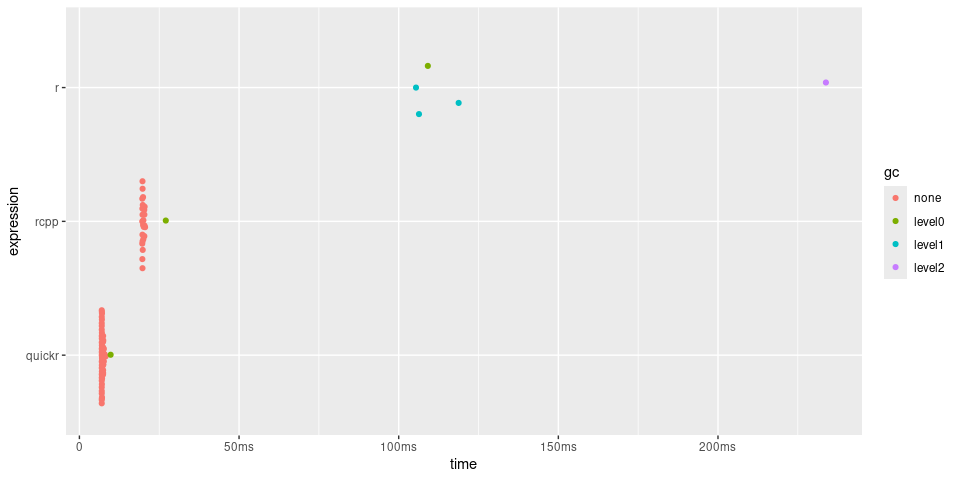
Using quickr in an R package
When called in a package, quick() will pre-compile the quick functions and place them in the ./src directory. Run devtools::load_all() or quickr::compile_package() to ensure that the generated files in ./src and ./R are in sync with each other.
Installation
You can install quickr from CRAN with:
install.packages("quickr")
You can install the development version of quickr from GitHub with:
# install.packages("pak")
pak::pak("t-kalinowski/quickr")
You will also need a C and Fortran compiler, preferably the same ones used to build R itself.
On macOS:
Make sure xcode tools and gfortran are installed, as described in https://mac.r-project.org/tools/. In Terminal, run:
sudo xcode-select --install # curl -LO https://mac.r-project.org/tools/gfortran-12.2-universal.pkg # R 4.4 curl -LO https://mac.r-project.org/tools/gfortran-14.2-universal.pkg # R 4.5 sudo installer -pkg gfortran-12.2-universal.pkg -target /
On Windows:
- Install the latest version of Rtools
On Linux:
- The “Install Required Dependencies” section here provides detailed instructions for installing R build tools on various Linux flavors.