Single-Source Capture-Recapture Models.
singleRcapture: An R Package for Single-Source Capture-Recapture Models 
Capture-recapture type experiments are used to estimate the total population size in situations when observing only a part of such population is feasible. In recent years these types of experiments have seen more interest.
Single-source models are distinct from other capture-recapture models because we cannot estimate the population size based on how many units were observed in two or three sources which is the standard approach.
Instead in single-source models we utilize count data regression models on positive distributions (i.e. on counts greater than 0) where the dependent variable is the number of times a particular unit was observed in source data.
This package aims to implement already existing and introduce new methods of estimating population size from single source to simplify the research process.
Currently, we have implemented most of the frequentist approaches used in literature such as:
- Zero-truncated Poisson, geometric and negative binomial regression.
- Zero-truncated one-inflated and one-inflated zero-truncated Poisson, geometric and negative binomial model.
- Zero-one-truncated Poisson, geometric and negative binomial models.
- Generalized Chao’s and Zelterman’s models based on logistic regression.
- Three types of bootstrap parametric, semi-parametric and nonparametric.
- And a wide range of additional functionalities associated with (vector) generalized linear models relevant to the topic.
For more details we see the singleRcapture: An R Package for Single-Source Capture-Recapture Models vignette on CRAN or pkgdown website.
Installation
You can install the current version of the singleRcapture package from main branch GitHub with:
# install.packages("devtools")
remotes::install_github("ncn-foreigners/singleRcapture")
or install the stable version from CRAN with:
install.packages(singleRcapture)
Examples
The main function of this package is estimatePopsize which fitts regression on specified distribution and then uses fitted regression to estimate the population size.
Lets look at a model from 2003 publication (Van Der Heijden, P. G., Bustami, R., Cruyff, M. J., Engbersen, G., & Van Houwelingen, H. C. (2003). Point and interval estimation of the population size using the truncated Poisson regression model. Statistical Modelling, 3(4), 305-322.). The call to estimatePopsize will look very similar to anyone who used the stats::glm function:
library(singleRcapture)
model <- estimatePopsize(
formula = capture ~ gender + age + nation, # specify formula
data = netherlandsimmigrant,
popVar = "analytic", # specify
model = "ztpoisson", # distribution used
method = "IRLS", # fitting method one of three currently supported
controlMethod = controlMethod(silent = TRUE) # ignore convergence at half step warning
)
summary(model) # a summary method for singleR class with standard glm-like output and population size estimation resutls
#>
#> Call:
#> estimatePopsize.default(formula = capture ~ gender + age + nation,
#> data = netherlandsimmigrant, model = "ztpoisson", method = "IRLS",
#> popVar = "analytic", controlMethod = controlMethod(silent = TRUE))
#>
#> Pearson Residuals:
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> -0.486442 -0.486442 -0.298080 0.002093 -0.209444 13.910844
#>
#> Coefficients:
#> -----------------------
#> For linear predictors associated with: lambda
#> Estimate Std. Error z value P(>|z|)
#> (Intercept) -1.3411 0.2149 -6.241 4.35e-10 ***
#> gendermale 0.3972 0.1630 2.436 0.014832 *
#> age>40yrs -0.9746 0.4082 -2.387 0.016972 *
#> nationAsia -1.0926 0.3016 -3.622 0.000292 ***
#> nationNorth Africa 0.1900 0.1940 0.979 0.327398
#> nationRest of Africa -0.9106 0.3008 -3.027 0.002468 **
#> nationSurinam -2.3364 1.0136 -2.305 0.021159 *
#> nationTurkey -1.6754 0.6028 -2.779 0.005445 **
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> AIC: 1712.901
#> BIC: 1757.213
#> Residual deviance: 1128.553
#>
#> Log-likelihood: -848.4504 on 1872 Degrees of freedom
#> Number of iterations: 8
#> -----------------------
#> Population size estimation results:
#> Point estimate 12690.35
#> Observed proportion: 14.8% (N obs = 1880)
#> Std. Error 2808.165
#> 95% CI for the population size:
#> lowerBound upperBound
#> normal 7186.449 18194.25
#> logNormal 8431.277 19718.31
#> 95% CI for the share of observed population:
#> lowerBound upperBound
#> normal 10.332933 26.16035
#> logNormal 9.534288 22.29793
We implemented a method for plot function to visualise the model fit and other useful diagnostic information. One of which is rootogram, a type of plot that compares fitted and observed marginal frequencies:
plot(model, plotType = "rootogram")
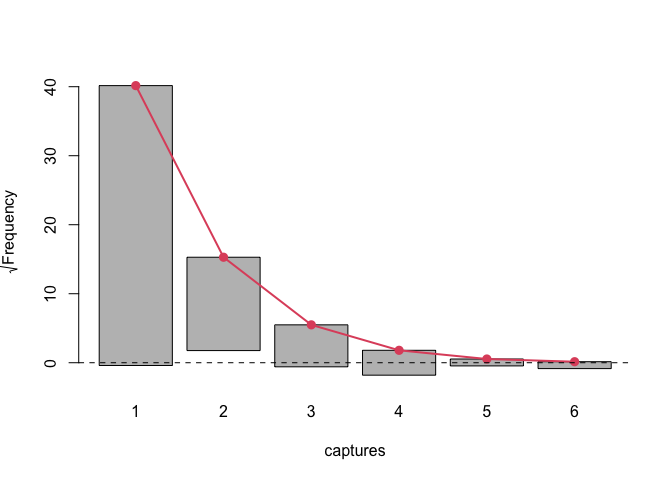
The possible values for plotType argument are:
qq- the normal quantile-quantile plot for pearson residuals (default),marginal- amatplotcomparing fitted and observed marginal frequencies,fitresid- plot of linear predictor values contrasted with pearson residuals,bootHist- histogram of bootstrap sample,rootogram- rootogram, example presented above,dfpopContr- contrasting two deletion effects to identify presence of influential observations,dfpopBox- boxplot of results fromdfpopsizefunction see its documentation,scaleLoc- scale-location plot,cooks- plot ofcooks.valuesfor distributions for which it is defined,hatplot- plot ofhatvalues,strata- plot of confidence intervals for selected such populations.
User can also pass arguments to specify additional information such as plot title, subtitle etc. similar to calling plot on some data. For more info check plot.singleR method documentation.
As we have seen there are some significant differences between fitted and observed marginal frequencies. To check our intuition let’s perform goodness of fit test between fitted and observed marginal frequencies.
To do it we call a summary function of marginalFreq function which computes marginal frequencies for the fitted singleR class object:
summary(marginalFreq(model), df = 2, dropl5 = "group")
#> Test for Goodness of fit of a regression model:
#>
#> Test statistics df P(>X^2)
#> Chi-squared test 50.06 2 1.3e-11
#> G-test 34.31 2 3.6e-08
#>
#> --------------------------------------------------------------
#> Cells with fitted frequencies of < 5 have been grouped
#> Names of cells used in calculating test(s) statistic: 1 2 3
Finally let us check if we have any influential observations. We will do this by comparing the deletion effect of every observation on population size estimate by removing it entirely from the model (from population size estimate and regression) and by only omitting it in pop size estimation (this is what is called the contribution of an observation). If observation is not influential these two actions should have the approximately the same effect:
plot(model, plotType = "dfpopContr")
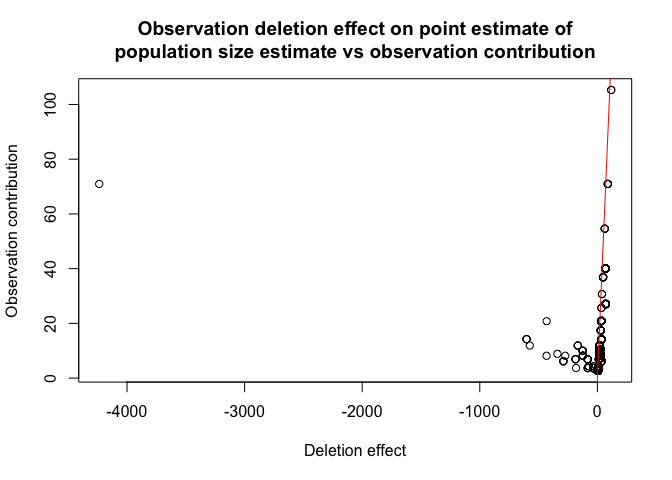
it is easy to deduce from the plot above that we have influential observations in our dataset (one in particular).
Lastly singleRcapture offers some post-hoc procedures for example a function stratifyPopsize that estimates sizes of user specified sub populations and returns them in a data.frame:
stratifyPopsize(model, alpha = c(.01, .02, .03, .05), # different significance level for each sub population
strata = list(
"Females from Surinam" = netherlandsimmigrant$gender == "female" & netherlandsimmigrant$nation == "Surinam",
"Males from Turkey" = netherlandsimmigrant$gender == "male" & netherlandsimmigrant$nation == "Turkey",
"Younger males" = netherlandsimmigrant$gender == "male" & netherlandsimmigrant$age == "<40yrs",
"Older males" = netherlandsimmigrant$gender == "male" & netherlandsimmigrant$age == ">40yrs"
))
#> name Observed Estimated ObservedPercentage StdError
#> 1 Females from Surinam 20 931.4677 2.147149 955.0657
#> 2 Males from Turkey 78 1291.2513 6.040652 741.0066
#> 3 Younger males 1391 7337.0708 18.958520 1282.1402
#> 4 Older males 91 1542.1886 5.900705 781.4747
#> normalLowerBound normalUpperBound logNormalLowerBound logNormalUpperBound
#> 1 -1528.61853 3391.554 119.2661 8389.158
#> 2 -432.58790 3015.090 405.4127 4573.791
#> 3 4554.71057 10119.431 5134.8122 10834.785
#> 4 10.52637 3073.851 630.7551 3992.674
#> confLevel
#> 1 0.01
#> 2 0.02
#> 3 0.03
#> 4 0.05
strata argument may be specified in various ways for example:
stratifyPopsize(model, strata = ~ gender / age)
#> name Observed Estimated ObservedPercentage StdError
#> 1 gender==female 398 3811.0911 10.443203 1153.9733
#> 2 gender==male 1482 8879.2594 16.690581 1812.0790
#> 3 genderfemale:age<40yrs 378 3169.8263 11.924944 880.9478
#> 4 gendermale:age<40yrs 1391 7337.0708 18.958520 1282.1402
#> 5 genderfemale:age>40yrs 20 641.2648 3.118836 407.5264
#> 6 gendermale:age>40yrs 91 1542.1886 5.900705 781.4747
#> normalLowerBound normalUpperBound logNormalLowerBound logNormalUpperBound
#> 1 1549.34513 6072.837 2189.0443 6902.133
#> 2 5327.64991 12430.869 6090.7762 13354.880
#> 3 1443.20030 4896.452 1904.3126 5484.617
#> 4 4824.12208 9850.019 5306.3306 10421.082
#> 5 -157.47223 1440.002 212.3382 2026.726
#> 6 10.52637 3073.851 630.7551 3992.674
#> confLevel
#> 1 0.05
#> 2 0.05
#> 3 0.05
#> 4 0.05
#> 5 0.05
#> 6 0.05
The package was designed with convenience in mind, for example it is possible to specify that weights provided on call are to be interpreted as number of occurrences of units in each row:
df <- netherlandsimmigrant[, c(1:3,5)]
df$ww <- 0
### this is dplyr::count without dependencies
df <- aggregate(ww ~ ., df, FUN = length)
summary(estimatePopsize(
formula = capture ~ nation + age + gender,
data = df,
model = ztpoisson,
weights = df$ww,
controlModel = controlModel(weightsAsCounts = TRUE)
))
#>
#> Call:
#> estimatePopsize.default(formula = capture ~ nation + age + gender,
#> data = df, model = ztpoisson, weights = df$ww, controlModel = controlModel(weightsAsCounts = TRUE))
#>
#> Pearson Residuals:
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> -317.6467 -2.4060 3.7702 0.0803 13.4920 183.2108
#>
#> Coefficients:
#> -----------------------
#> For linear predictors associated with: lambda
#> Estimate Std. Error z value P(>|z|)
#> (Intercept) -1.3411 0.2149 -6.241 4.35e-10 ***
#> nationAsia -1.0926 0.3016 -3.622 0.000292 ***
#> nationNorth Africa 0.1900 0.1940 0.979 0.327398
#> nationRest of Africa -0.9106 0.3008 -3.027 0.002468 **
#> nationSurinam -2.3364 1.0136 -2.305 0.021159 *
#> nationTurkey -1.6754 0.6028 -2.779 0.005445 **
#> age>40yrs -0.9746 0.4082 -2.387 0.016972 *
#> gendermale 0.3972 0.1630 2.436 0.014832 *
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> AIC: 1712.901
#> BIC: 1757.213
#> Residual deviance: 1128.553
#>
#> Log-likelihood: -848.4504 on 1872 Degrees of freedom
#> Number of iterations: 8
#> -----------------------
#> Population size estimation results:
#> Point estimate 12690.35
#> Observed proportion: 14.8% (N obs = 1880)
#> Std. Error 2808.169
#> 95% CI for the population size:
#> lowerBound upperBound
#> normal 7186.444 18194.26
#> logNormal 8431.275 19718.32
#> 95% CI for the share of observed population:
#> lowerBound upperBound
#> normal 10.332927 26.16037
#> logNormal 9.534281 22.29793
Methods such as regression diagnostics will be adjusted (values of weights will be reduced instead of rows being removed etc.)
We also included option to use common non standard argument such as significance levels different from usual 5%:
set.seed(123)
modelInflated <- estimatePopsize(
formula = capture ~ gender + age,
data = netherlandsimmigrant,
model = "ztoigeom",
method = "IRLS",
# control parameters for population size estimation check documentation of controlPopVar
controlPopVar = controlPopVar(
alpha = .01, # significance level
)
)
summary(modelInflated)
#>
#> Call:
#> estimatePopsize.default(formula = capture ~ gender + age, data = netherlandsimmigrant,
#> model = "ztoigeom", method = "IRLS", controlPopVar = controlPopVar(alpha = 0.01,
#> ))
#>
#> Pearson Residuals:
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> -0.357193 -0.357193 -0.357193 0.000343 -0.287637 10.233608
#>
#> Coefficients:
#> -----------------------
#> For linear predictors associated with: lambda
#> Estimate Std. Error z value P(>|z|)
#> (Intercept) -1.5346 0.1846 -8.312 < 2e-16 ***
#> gendermale 0.3863 0.1380 2.800 0.00512 **
#> age>40yrs -0.7788 0.2942 -2.648 0.00810 **
#> -----------------------
#> For linear predictors associated with: omega
#> Estimate Std. Error z value P(>|z|)
#> (Intercept) -1.7591 0.3765 -4.673 2.97e-06 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> AIC: 1736.854
#> BIC: 1759.01
#> Residual deviance: 1011.271
#>
#> Log-likelihood: -864.4272 on 3756 Degrees of freedom
#> Number of iterations: 6
#> -----------------------
#> Population size estimation results:
#> Point estimate 5661.522
#> Observed proportion: 33.2% (N obs = 1880)
#> Std. Error 963.9024
#> 99% CI for the population size:
#> lowerBound upperBound
#> normal 3178.674 8144.370
#> logNormal 3861.508 9096.681
#> 99% CI for the share of observed population:
#> lowerBound upperBound
#> normal 23.08343 59.14416
#> logNormal 20.66688 48.68564
and the option to estimate standard error of population size estimate by bootstrap, models with more than one distribution parameter being dependent on covariates and some non standard link functions for example:
modelInflated2 <- estimatePopsize(
formula = capture ~ age,
data = netherlandsimmigrant,
popVar = "bootstrap",
model = ztoigeom(omegaLink = "cloglog"),
method = "IRLS",
controlPopVar = controlPopVar(
B = 500,# number of boostrap samples
alpha = .01, # significance level
# type of bootstrap see documentation for estimatePopsize
bootType = "semiparametric",
# control regression fitting on bootstrap samples
bootstrapFitcontrol = controlMethod(
epsilon = .Machine$double.eps,
silent = TRUE,
stepsize = 2
)
),
controlModel = controlModel(omegaFormula = ~ gender) # put covariates on omega i.e. the inflation parameter
)
#> Warning in estimatePopsize.default(formula = capture ~ age, data = netherlandsimmigrant, : The (analytically computed) hessian of the score function is not negative define.
#> NOTE: Second derivative test failing does not
#> necessarily mean that the maximum of score function that was found
#> numericaly is invalid since R^k is not a bounded space.
#> Additionally in one inflated and hurdle models second derivative test often fails even on valid arguments.
#> Warning in estimatePopsize.default(formula = capture ~ age, data =
#> netherlandsimmigrant, : Switching from observed information matrix to Fisher
#> information matrix because hessian of log-likelihood is not negative define.
popSizeEst(modelInflated2)
#> Point estimate: 5496.376
#> Variance: 1217986
#> 99% confidence intervals:
#> lowerBound upperBound
#> 4002.763 10231.716
the results are significantly different (the warning issued concerns the second derivative test for existence of local minimum, here it was inconclusive but we manually checked that fitting process found the optimal regression coefficients it’s here to provide more information to the user):
plot(modelInflated2,
plotType = "bootHist",
labels = TRUE,
ylim = c(0, 175),
breaks = 15)
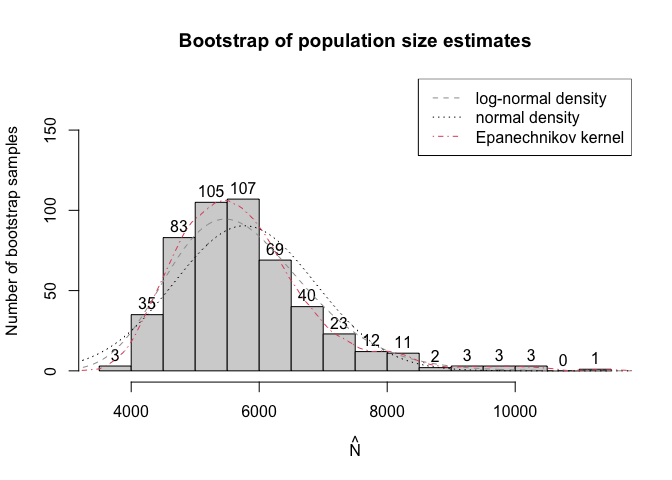
and information criteria support the second model:
#> First model: AIC = 1736.854 BIC = 1759.01
#> Second model: AIC = 1734.803 BIC = 1756.959
Funding
Work on this package is supported by the the National Science Centre, OPUS 20 grant no. 2020/39/B/HS4/00941.