Optimal Histogram Binning Using Shimazaki-Shinomoto Method.
sshist
The sshist package implements the Shimazaki-Shinomoto method for finding the optimal number of bins in histograms.
Unlike the standard Freedman-Diaconis rule (used by default in ggplot2), this method minimizes the expected L2 loss function between the histogram and the unknown underlying density function. It is particularly effective for:
- Time-dependent rate estimation (PSTH).
- Identifying intrinsic data structures in multimodal distributions.
- Optimizing both 1D and 2D data binnings.
Installation
# stable version from CRAN
install.packages("sshist")
You can install the development version of sshist like so:
# install.packages("devtools")
devtools::install_github("celebithil/sshist")
Example 1: Basic 1D Usage
Here is a basic example using the Old Faithful Geyser data.
library(sshist)
# Load data
data(faithful)
x_data <- faithful$waiting
# Calculate optimal binning
res <- sshist(x_data)
# Print summary
print(res)
#> Shimazaki-Shinomoto Histogram Optimization
#> ------------------------------------------
#> Optimal Bins (N): 21
#> Bin Width (D): 2.524
#> Cost Minimum: -8.525
hist(res$data, breaks=res$edges, freq=FALSE,
main=paste("Optimal Hist (N=", res$opt_n, ")"),
col="lightblue", border="white", xlab="Data")
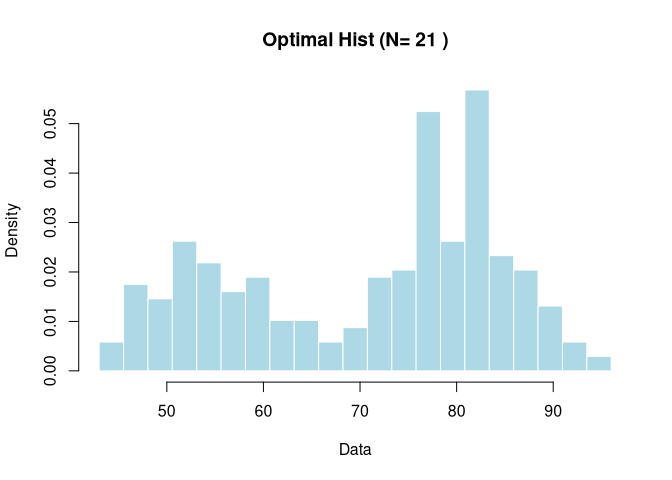
Example 2: Integration with ggplot2
sshist calculates the optimal parameters, which you can easily pass to ggplot2.
library(ggplot2)
# Create a data frame
df <- data.frame(waiting = x_data)
ggplot(df, aes(x = waiting)) +
geom_histogram(breaks = res$edges, fill = "#69b3a2", color = "white", alpha = 0.8) +
geom_rug(alpha = 0.1) +
ggtitle(paste0("Shimazaki-Shinomoto Optimization (N = ", res$opt_n, ")")) +
theme_minimal()
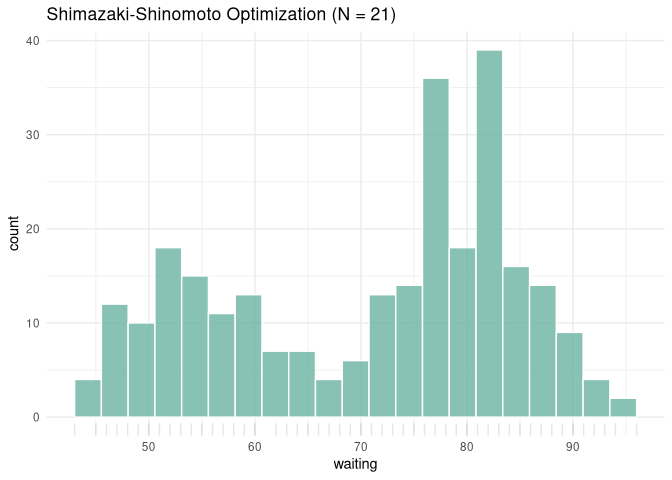
Example 3: 2D Histogram Optimization
For bivariate data, sshist_2d finds the optimal binning for both X and Y axes simultaneously.
# Get bimodal 2D data
y_data <- faithful$eruptions
# Optimize
res2d <- sshist_2d(x_data, y_data)
# Print summary
print(res2d)
#> Shimazaki-Shinomoto 2D Histogram Optimization
#> ---------------------------------------------
#> Optimal Bins X: 9
#> Optimal Bins Y: 20
#> Bin Width X: 5.889
#> Bin Width Y: 0.175
#> Cost Minimum: -5.717
Example 4: 2D Optimization with ggplot2
You can easily use the optimized bin counts from sshist_2d in ggplot2 by passing them to the bins argument in geom_bin2d.
# We use the 'res2d' object calculated in Example 3
# containing optimal bins for Old Faithful data
res2d <- sshist_2d(faithful$waiting, faithful$eruptions )
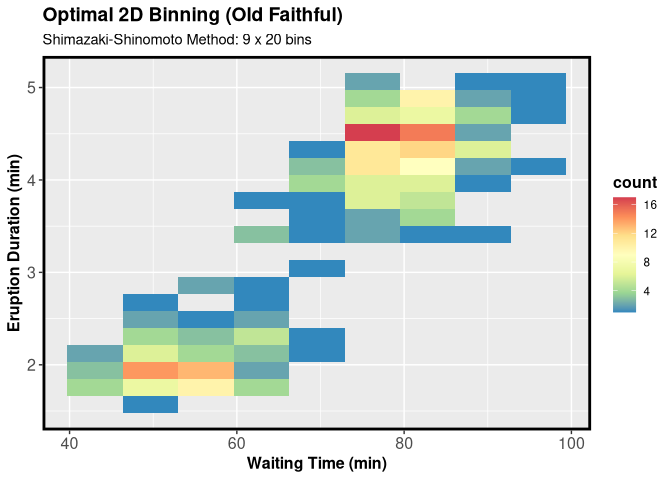
ggplot(faithful, aes(waiting, eruptions)) +
geom_bin2d(bins = c(res2d$opt_nx, res2d$opt_ny)) +
scale_fill_distiller(palette = "Spectral") +
labs(
title = "Optimal 2D Binning (Old Faithful)",
subtitle = paste0("Shimazaki-Shinomoto Method: ",
res2d$opt_nx, " x ", res2d$opt_ny, " bins"),
x = "Waiting Time (min)",
y = "Eruption Duration (min)"
) +
theme(axis.text = element_text(size = 12),
title = element_text(size = 12,face="bold"),
panel.border = element_rect(linewidth = 2, color = "black", fill = NA))
References
- Shimazaki, H. and Shinomoto, S., 2007. A method for selecting the bin size of a time histogram. Neural Computation, 19(6), pp.1503-1527. doi:10.1162/neco.2007.19.6.1503
- https://www.neuralengine.org/res/histogram.html
- https://github.com/shimazaki/density_estimation
- https://s-shinomoto.com/toolbox/sshist/hist.html.