Description
Collapsed Variational Inference for Dirichlet Process (DP) Mixture Model.
Description
Collapsed Variational Inference for a Dirichlet Process (DP) mixture model with unknown covariance matrix structure and DP concentration parameter. It enables efficient clustering of high-dimensional data with significantly improved computational speed than traditional MCMC methods. The package incorporates 8 parameterisations and corresponding prior choices for the unknown covariance matrix, from which the user can choose and apply accordingly.
README.md
vimixr
The goal of vimixr is to perform collapsed Variational Inference for DPMM using adaptive inference on the DP concentration parameter as well as covariance hyper-parameter of DP base distribution.
Installation
You can install the development version of vimixr from GitHub with:
# install.packages("devtools")
devtools::install_github("annesh07/vimixr")
Example
library(vimixr)
Let’s generate some toy data. Here the data contains N = 100 samples, D = 2 dimensions and K = 2 clusters.
X <- rbind(matrix(rnorm(100, m=0, sd=0.5), ncol=2),
matrix(rnorm(100, m=3, sd=0.5), ncol=2))
In order to obtain the clusters present in X, we apply function from vimixr package that corresponds to a simple case with cluster-inspecific fixed diagonal covariance for the data.
# Fixed-diagonal variance
res <- cvi_npmm(X, variational_params = 20, prior_shape_alpha = 0.001,
prior_rate_alpha = 0.001, post_shape_alpha = 0.001,
post_rate_alpha = 0.001, prior_mean_eta = matrix(0, 1, ncol(X)),
post_mean_eta = matrix(0.001, 20, ncol(X)),
log_prob_matrix = t(apply(matrix(0.001, nrow(X), 20), 1,
function(x){x/sum(x)})), maxit = 1000,
fixed_variance = TRUE, covariance_type = "diagonal",
prior_precision_scalar_eta = 0.001,
post_precision_scalar_eta = matrix(0.001, 20, 1),
cov_data = diag(ncol(X)))
summary(res)
#> Length Class Mode
#> posterior 5 -none- list
#> optimisation 3 -none- list
#> PCA_viz 1 ggplot2::ggplot S4
#> ELBO_viz 1 ggplot2::ggplot S4
#> Seed_used 1 -none- character
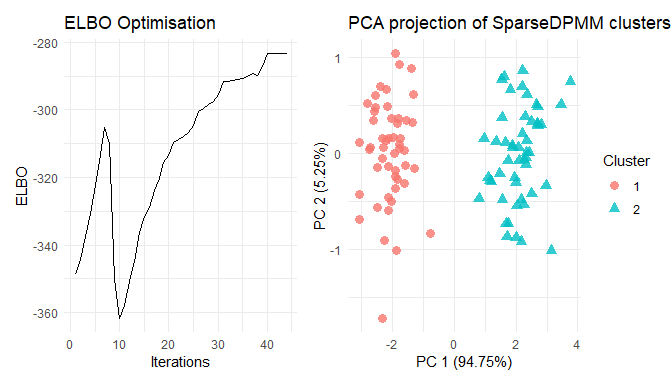
plot(res)